You are browsing environment: FUNGIDB
CAZyme Information: ASPTUDRAFT_172437-t33_1-p1
You are here: Home > Sequence: ASPTUDRAFT_172437-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tubingensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tubingensis | |||||||||||
| CAZyme ID | ASPTUDRAFT_172437-t33_1-p1 | |||||||||||
| CAZy Family | GH128 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 33 | 328 | 5.3e-76 | 0.976897689768977 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 214750 | Glyco_10 | 7.25e-85 | 74 | 327 | 1 | 262 | Glycosyl hydrolase family 10. |
| 395262 | Glyco_hydro_10 | 5.67e-78 | 31 | 327 | 1 | 307 | Glycosyl hydrolase family 10. |
| 226217 | XynA | 2.77e-67 | 21 | 327 | 14 | 336 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.25e-185 | 1 | 335 | 1 | 336 | |
| 1.19e-182 | 1 | 335 | 1 | 331 | |
| 5.07e-180 | 1 | 342 | 1 | 343 | |
| 3.34e-167 | 1 | 337 | 1 | 338 | |
| 1.30e-163 | 9 | 335 | 6 | 332 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.08e-48 | 31 | 332 | 4 | 302 | Crystal Structure of the catalytic domain of xylanase A from Streptomyces halstedii JM8 [Streptomyces halstedii] |
|
| 3.76e-45 | 31 | 332 | 6 | 320 | Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBR_B Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBU_A Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima],1VBU_B Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima] |
|
| 5.46e-44 | 31 | 332 | 22 | 336 | Crystal structure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NIY_B Crystal structure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NJ3_A Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1],3NJ3_B Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1] |
|
| 8.79e-44 | 31 | 335 | 28 | 347 | Chain A, 1,4-BETA-D-XYLAN-XYLANOHYDROLASE [Acetivibrio thermocellus],1XYZ_B Chain B, 1,4-BETA-D-XYLAN-XYLANOHYDROLASE [Acetivibrio thermocellus] |
|
| 2.81e-43 | 46 | 327 | 26 | 337 | Highly active enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.46e-41 | 60 | 333 | 58 | 327 | Endo-1,4-beta-xylanase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xlnC PE=1 SV=1 |
|
| 1.20e-40 | 31 | 335 | 518 | 837 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
|
| 1.54e-40 | 50 | 326 | 50 | 347 | Beta-1,4-xylanase OS=Dictyoglomus thermophilum OX=14 GN=xynA PE=3 SV=1 |
|
| 1.67e-40 | 52 | 332 | 52 | 326 | Probable endo-1,4-beta-xylanase C OS=Aspergillus terreus OX=33178 GN=xlnC PE=2 SV=1 |
|
| 1.67e-40 | 52 | 332 | 52 | 326 | Probable endo-1,4-beta-xylanase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=xlnC PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
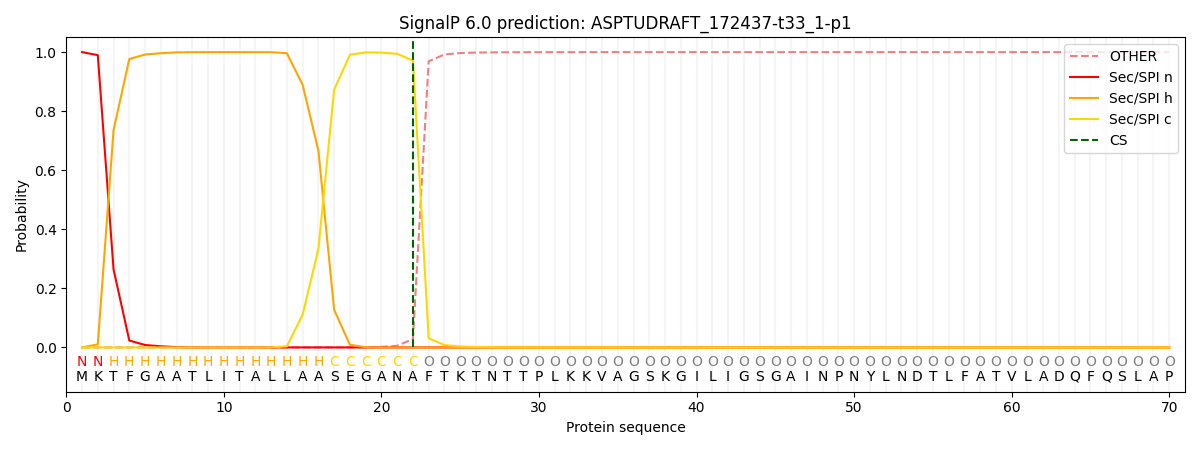
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000236 | 0.999743 | CS pos: 22-23. Pr: 0.9715 |
