You are browsing environment: FUNGIDB
CAZyme Information: ASPTUDRAFT_125250-t33_1-p1
You are here: Home > Sequence: ASPTUDRAFT_125250-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tubingensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tubingensis | |||||||||||
| CAZyme ID | ASPTUDRAFT_125250-t33_1-p1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH17 | 397 | 669 | 2e-33 | 0.9485530546623794 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 3.82e-39 | 399 | 666 | 60 | 305 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 366033 | Glyco_hydro_17 | 5.21e-14 | 403 | 669 | 25 | 304 | Glycosyl hydrolases family 17. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 683 | 1 | 681 | |
| 0.0 | 1 | 683 | 1 | 681 | |
| 0.0 | 1 | 683 | 1 | 662 | |
| 3.26e-290 | 1 | 681 | 1 | 683 | |
| 6.56e-290 | 1 | 681 | 1 | 683 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.12e-33 | 385 | 674 | 36 | 294 | Crystal structure of glycoside hydrolase family 17 beta-1,3-glucanosyltransferase from Rhizomucor miehei [Rhizomucor miehei CAU432] |
|
| 5.19e-32 | 385 | 674 | 36 | 294 | Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaribiose [Rhizomucor miehei CAU432],4WTS_A Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaritriose [Rhizomucor miehei CAU432] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 683 | 1 | 684 | Putative glucan endo-1,3-beta-glucosidase btgC OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=btgC PE=3 SV=2 |
|
| 1.17e-290 | 1 | 681 | 1 | 683 | Probable glucan endo-1,3-beta-glucosidase btgC OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=btgC PE=3 SV=1 |
|
| 1.17e-290 | 1 | 681 | 1 | 683 | Probable glucan endo-1,3-beta-glucosidase btgC OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=btgC PE=3 SV=2 |
|
| 6.57e-274 | 1 | 683 | 1 | 688 | Probable glucan endo-1,3-beta-glucosidase btgC OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=btgC PE=3 SV=1 |
|
| 1.81e-273 | 1 | 683 | 1 | 688 | Probable glucan endo-1,3-beta-glucosidase btgC OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=btgC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
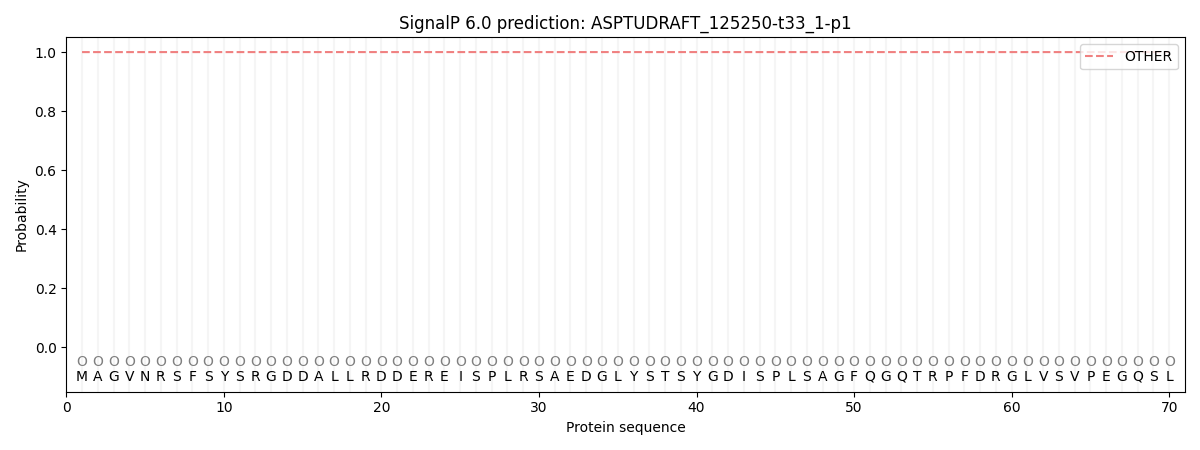
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000050 | 0.000000 |

