You are browsing environment: FUNGIDB
CAZyme Information: ASPSYDRAFT_43114-t33_1-p1
You are here: Home > Sequence: ASPSYDRAFT_43114-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus sydowii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus sydowii | |||||||||||
| CAZyme ID | ASPSYDRAFT_43114-t33_1-p1 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT34 | 146 | 256 | 2.7e-20 | 0.43902439024390244 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 368535 | Glyco_transf_34 | 1.29e-23 | 60 | 254 | 21 | 231 | galactosyl transferase GMA12/MNN10 family. This family contains a number of glycosyltransferase enzymes that contain a DXD motif. This family includes a number of C. elegans homologs where the DXD is replaced by DXH. Some members of this family are included in glycosyltransferase family 34. |
| 215616 | PLN03181 | 6.62e-08 | 74 | 207 | 159 | 309 | glycosyltransferase; Provisional |
| 215617 | PLN03182 | 4.14e-07 | 44 | 178 | 115 | 264 | xyloglucan 6-xylosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.44e-219 | 1 | 318 | 65 | 382 | |
| 6.52e-161 | 1 | 314 | 14 | 325 | |
| 2.07e-121 | 35 | 297 | 283 | 549 | |
| 2.42e-119 | 17 | 295 | 2 | 284 | |
| 1.15e-97 | 57 | 310 | 1 | 243 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.28e-11 | 54 | 242 | 121 | 317 | Probable glycosyltransferase 7 OS=Oryza sativa subsp. indica OX=39946 GN=GT7 PE=3 SV=2 |
|
| 1.28e-11 | 54 | 242 | 121 | 317 | Probable glycosyltransferase 7 OS=Oryza sativa subsp. japonica OX=39947 GN=GT7 PE=2 SV=1 |
|
| 3.55e-10 | 62 | 284 | 121 | 364 | Uncharacterized alpha-1,2-galactosyltransferase C1289.13c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC1289.13c PE=3 SV=1 |
|
| 1.54e-09 | 48 | 284 | 97 | 364 | Alpha-1,2-galactosyltransferase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gma12 PE=1 SV=1 |
|
| 2.31e-09 | 90 | 259 | 132 | 315 | Probable alpha-1,2-galactosyltransferase gmh1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gmh1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
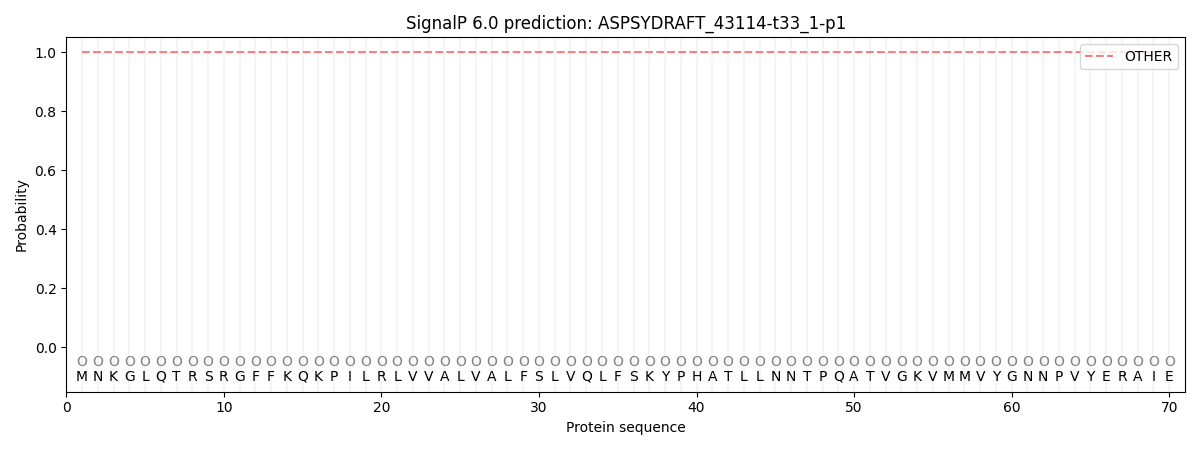
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000001 | 0.000028 |

