You are browsing environment: FUNGIDB
CAZyme Information: ASPCADRAFT_150423-t33_1-p1
You are here: Home > Sequence: ASPCADRAFT_150423-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus carbonarius | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus carbonarius | |||||||||||
| CAZyme ID | ASPCADRAFT_150423-t33_1-p1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | glycoside hydrolase family 55 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.58:16 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 37 | 807 | 4.4e-240 | 0.9891891891891892 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 403800 | Pectate_lyase_3 | 4.29e-80 | 65 | 307 | 1 | 213 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 403800 | Pectate_lyase_3 | 4.05e-06 | 435 | 489 | 4 | 62 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 227721 | Pgu1 | 8.52e-06 | 67 | 124 | 84 | 125 | Polygalacturonase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 968 | 46 | 1050 | |
| 0.0 | 1 | 962 | 1 | 949 | |
| 0.0 | 1 | 862 | 1 | 891 | |
| 1.39e-256 | 39 | 815 | 40 | 804 | |
| 7.71e-256 | 1 | 863 | 45 | 917 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.04e-193 | 39 | 807 | 21 | 755 | Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQN_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_A Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium] |
|
| 1.37e-180 | 44 | 805 | 6 | 751 | Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila],5M60_A Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila] |
|
| 1.05e-06 | 67 | 129 | 76 | 128 | Chain A, Putative pectin lyase [Geobacillus virus E2],7CHU_B Chain B, Putative pectin lyase [Geobacillus virus E2],7CHU_C Chain C, Putative pectin lyase [Geobacillus virus E2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.95e-192 | 41 | 805 | 40 | 871 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
|
| 1.35e-152 | 27 | 748 | 28 | 721 | Glucan 1,3-beta-glucosidase OS=Cochliobolus carbonum OX=5017 GN=EXG1 PE=1 SV=1 |
|
| 9.00e-42 | 110 | 801 | 94 | 747 | Glucan endo-1,3-beta-glucosidase BGN13.1 OS=Trichoderma harzianum OX=5544 GN=bgn13.1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
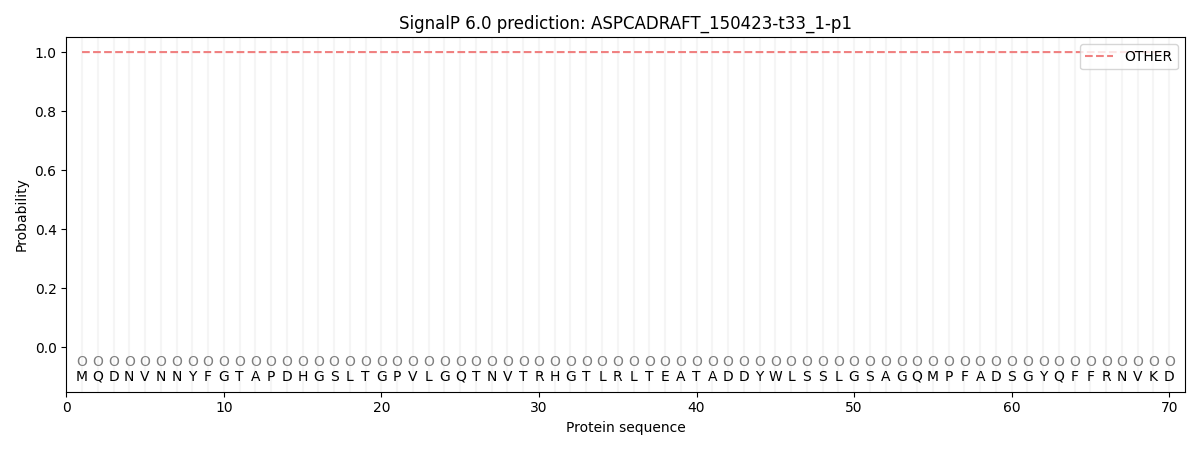
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000073 | 0.000000 |
