You are browsing environment: FUNGIDB
CAZyme Information: ASPBRDRAFT_193148-t33_1-p1
You are here: Home > Sequence: ASPBRDRAFT_193148-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus brasiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus brasiliensis | |||||||||||
| CAZyme ID | ASPBRDRAFT_193148-t33_1-p1 | |||||||||||
| CAZy Family | GH13|GH13 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.15:80 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 48 | 368 | 8.7e-75 | 0.9569230769230769 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395231 | Glyco_hydro_28 | 3.17e-148 | 52 | 368 | 3 | 321 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| 177865 | PLN02218 | 7.15e-27 | 87 | 362 | 143 | 420 | polygalacturonase ADPG |
| 215540 | PLN03010 | 1.07e-26 | 88 | 367 | 122 | 394 | polygalacturonase |
| 227721 | Pgu1 | 1.91e-24 | 72 | 295 | 174 | 408 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| 215426 | PLN02793 | 2.48e-22 | 104 | 359 | 145 | 406 | Probable polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.25e-256 | 1 | 368 | 1 | 368 | |
| 5.25e-256 | 1 | 368 | 1 | 368 | |
| 1.76e-254 | 1 | 368 | 1 | 368 | |
| 1.76e-254 | 1 | 368 | 1 | 368 | |
| 6.89e-238 | 1 | 367 | 1 | 367 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.96e-235 | 33 | 368 | 1 | 336 | Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_B Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_C Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_D Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_E Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_F Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger] |
|
| 5.33e-163 | 30 | 368 | 6 | 344 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 3.07e-162 | 30 | 368 | 6 | 344 | Chain A, endo-polygalacturonase [Evansstolkia leycettana] |
|
| 5.39e-161 | 35 | 368 | 3 | 336 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 9.63e-153 | 35 | 366 | 3 | 335 | Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_B Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_C Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_D Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_E Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_F Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_G Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.12e-255 | 1 | 368 | 1 | 368 | Endopolygalacturonase I OS=Aspergillus niger OX=5061 GN=pgaI PE=1 SV=1 |
|
| 3.12e-255 | 1 | 368 | 1 | 368 | Probable endopolygalacturonase I OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=pgaI PE=3 SV=1 |
|
| 3.81e-217 | 1 | 368 | 1 | 368 | Probable endopolygalacturonase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=pgaA PE=3 SV=1 |
|
| 1.27e-215 | 1 | 368 | 1 | 368 | Probable endopolygalacturonase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=pgaA PE=3 SV=1 |
|
| 1.27e-215 | 1 | 368 | 1 | 368 | Probable endopolygalacturonase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pgaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
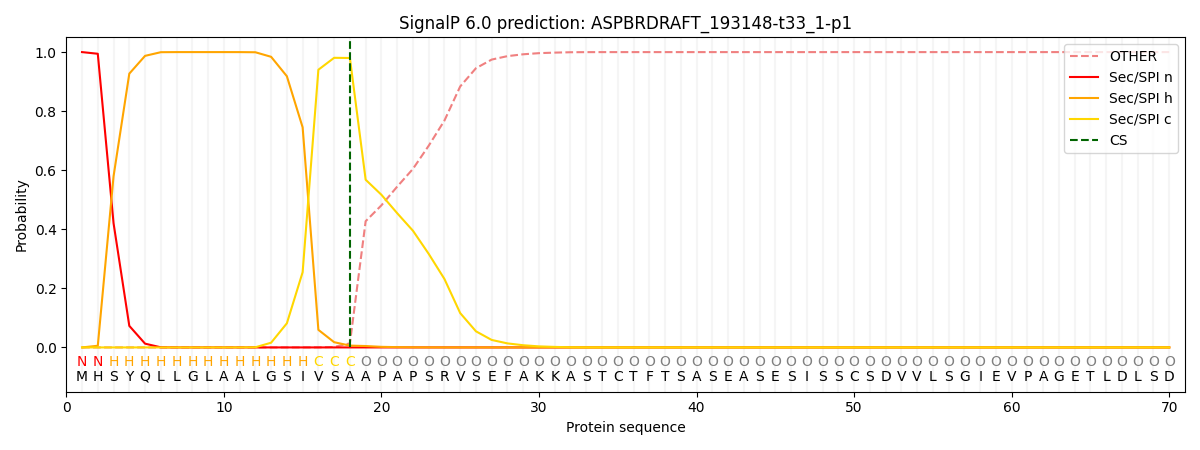
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000261 | 0.999735 | CS pos: 18-19. Pr: 0.9801 |
