You are browsing environment: FUNGIDB
CAZyme Information: ASPACDRAFT_74440-t33_1-p1
You are here: Home > Sequence: ASPACDRAFT_74440-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus aculeatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus aculeatus | |||||||||||
| CAZyme ID | ASPACDRAFT_74440-t33_1-p1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.22:58 | 2.4.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH27 | 125 | 418 | 2.5e-66 | 0.9912663755458515 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 269893 | GH27 | 4.24e-121 | 28 | 331 | 1 | 271 | glycosyl hydrolase family 27 (GH27). GH27 enzymes occur in eukaryotes, prokaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-N-acetylgalactosaminidase, and 3-alpha-isomalto-dextranase. All GH27 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH27 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| 166449 | PLN02808 | 7.61e-92 | 1 | 431 | 8 | 376 | alpha-galactosidase |
| 177874 | PLN02229 | 2.38e-90 | 24 | 391 | 59 | 376 | alpha-galactosidase |
| 178295 | PLN02692 | 6.20e-84 | 22 | 391 | 50 | 367 | alpha-galactosidase |
| 374582 | Melibiase_2 | 3.20e-68 | 28 | 331 | 2 | 284 | Alpha galactosidase A. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.62e-269 | 15 | 442 | 14 | 442 | |
| 1.62e-269 | 15 | 442 | 14 | 442 | |
| 1.38e-268 | 6 | 442 | 5 | 443 | |
| 1.38e-268 | 6 | 442 | 5 | 443 | |
| 1.61e-267 | 6 | 442 | 5 | 443 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.79e-174 | 18 | 442 | 1 | 414 | Chain A, alpha-galactosidase [Trichoderma reesei],1T0O_A Chain A, alpha-galactosidase [Trichoderma reesei] |
|
| 2.38e-84 | 22 | 438 | 3 | 360 | Nicotiana benthamiana alpha-galactosidase [Nicotiana benthamiana] |
|
| 7.25e-83 | 21 | 438 | 2 | 359 | Chain A, alpha-galactosidase [Oryza sativa] |
|
| 1.55e-78 | 28 | 441 | 9 | 393 | Crystal structure of alpha-galactosidase I from Mortierella vinacea [Umbelopsis vinacea] |
|
| 2.39e-71 | 22 | 375 | 24 | 359 | Chain A, Alpha-galactosidase 1 [Saccharomyces cerevisiae],3LRL_A Chain A, Alpha-galactosidase 1 [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.45e-269 | 6 | 442 | 5 | 443 | Probable alpha-galactosidase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=aglB PE=3 SV=1 |
|
| 2.86e-268 | 6 | 442 | 5 | 443 | Probable alpha-galactosidase B OS=Aspergillus niger OX=5061 GN=aglB PE=2 SV=1 |
|
| 3.09e-258 | 9 | 442 | 11 | 442 | Probable alpha-galactosidase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=aglB PE=3 SV=1 |
|
| 4.38e-258 | 9 | 442 | 11 | 442 | Probable alpha-galactosidase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=aglB PE=3 SV=1 |
|
| 4.79e-254 | 2 | 442 | 7 | 447 | Probable alpha-galactosidase B OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=aglB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
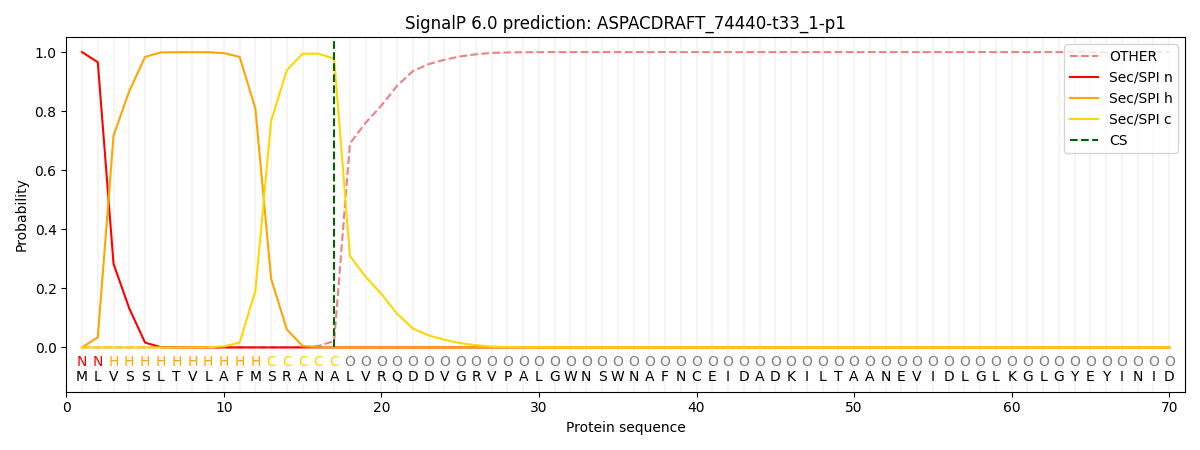
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000292 | 0.999686 | CS pos: 17-18. Pr: 0.9781 |
