You are browsing environment: FUNGIDB
CAZyme Information: APA15870.1
You are here: Home > Sequence: APA15870.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sclerotinia sclerotiorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Leotiomycetes; ; Sclerotiniaceae; Sclerotinia; Sclerotinia sclerotiorum | |||||||||||
| CAZyme ID | APA15870.1 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH12 | 115 | 258 | 1.9e-26 | 0.8974358974358975 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396303 | Glyco_hydro_12 | 2.24e-13 | 50 | 258 | 2 | 194 | Glycosyl hydrolase family 12. |
| 235746 | PRK06215 | 4.18e-07 | 2 | 201 | 8 | 171 | hypothetical protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.21e-200 | 1 | 274 | 1 | 274 | |
| 2.00e-171 | 1 | 274 | 1 | 274 | |
| 5.71e-171 | 1 | 274 | 1 | 274 | |
| 2.81e-158 | 1 | 274 | 1 | 274 | |
| 5.54e-138 | 10 | 270 | 7 | 266 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.35e-19 | 26 | 257 | 3 | 215 | Crystal structure of XEG [Aspergillus aculeatus],3VL9_A Crystal structure of xeg-xyloglucan [Aspergillus aculeatus],3VL9_B Crystal structure of xeg-xyloglucan [Aspergillus aculeatus] |
|
| 2.01e-17 | 35 | 257 | 5 | 208 | Crystal structure of xeg-edgp [Aspergillus aculeatus],3VLB_D Crystal structure of xeg-edgp [Aspergillus aculeatus] |
|
| 3.18e-14 | 37 | 256 | 3 | 204 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus],5GM3_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus] |
|
| 1.14e-13 | 37 | 228 | 3 | 175 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_C Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_D Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_E Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_F Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_G Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus] |
|
| 1.16e-13 | 37 | 228 | 4 | 176 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_C Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_D Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_E Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_F Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_G Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.02e-20 | 27 | 257 | 14 | 227 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus niger OX=5061 GN=xgeA PE=1 SV=1 |
|
| 2.14e-19 | 27 | 257 | 14 | 227 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=xgeA PE=3 SV=1 |
|
| 7.65e-19 | 27 | 257 | 13 | 225 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xgeA PE=1 SV=1 |
|
| 1.41e-17 | 27 | 257 | 13 | 224 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus aculeatus OX=5053 GN=xgeA PE=1 SV=1 |
|
| 1.44e-16 | 28 | 263 | 13 | 234 | Xyloglucan-specific endo-beta-1,4-glucanase 1 OS=Phytophthora sojae (strain P6497) OX=1094619 GN=XEG1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
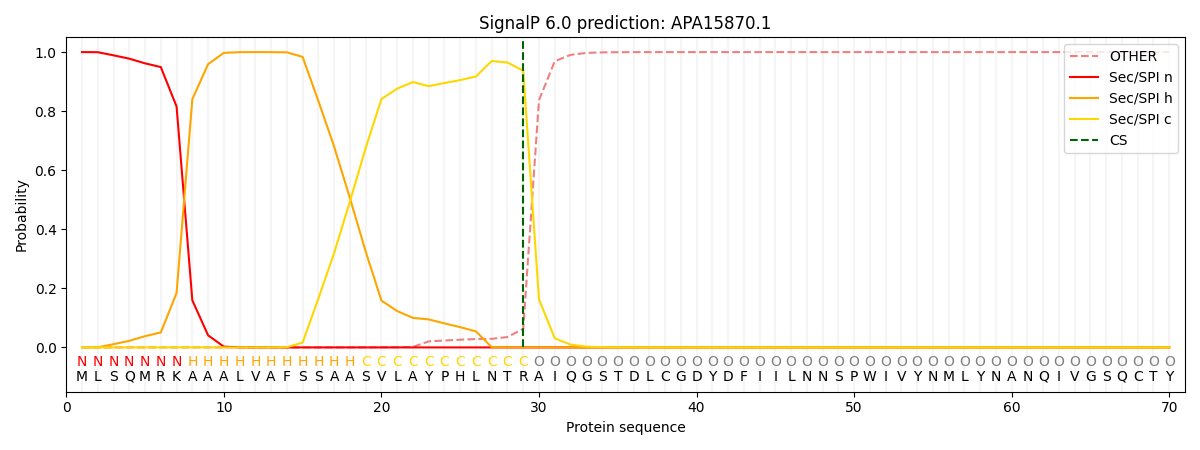
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000301 | 0.999695 | CS pos: 29-30. Pr: 0.9371 |
