You are browsing environment: FUNGIDB
CAZyme Information: APA06883.1
You are here: Home > Sequence: APA06883.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sclerotinia sclerotiorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Leotiomycetes; ; Sclerotiniaceae; Sclerotinia; Sclerotinia sclerotiorum | |||||||||||
| CAZyme ID | APA06883.1 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 50 | 521 | 1.6e-126 | 0.9977578475336323 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 0.0 | 50 | 521 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.75e-112 | 22 | 525 | 52 | 522 | glycoside hydrolase family 47 protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 526 | 1 | 526 | |
| 0.0 | 1 | 526 | 1 | 526 | |
| 0.0 | 1 | 526 | 1 | 557 | |
| 0.0 | 1 | 526 | 1 | 530 | |
| 0.0 | 1 | 526 | 1 | 530 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.42e-214 | 46 | 522 | 8 | 474 | Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI8_B Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI9_A Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum],2RI9_B Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum] |
|
| 1.95e-213 | 46 | 522 | 43 | 509 | Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KKT_B Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KRE_A Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRE_B Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_A Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_B Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum] |
|
| 3.85e-184 | 34 | 521 | 9 | 492 | Chain A, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_B Chain B, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_C Chain C, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_D Chain D, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei] |
|
| 2.66e-62 | 38 | 521 | 17 | 467 | Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus],5KKB_B Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus] |
|
| 3.29e-62 | 38 | 521 | 15 | 465 | Structure of mouse Golgi alpha-1,2-mannosidase IA reveals the molecular basis for substrate specificity among Class I enzymes (family 47 glycosidases) [Mus musculus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.99e-222 | 8 | 521 | 3 | 507 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mns1B PE=3 SV=1 |
|
| 4.90e-218 | 40 | 521 | 32 | 508 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=mns1B PE=3 SV=1 |
|
| 5.26e-218 | 1 | 522 | 3 | 511 | Mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus phoenicis OX=5063 GN=mns1B PE=2 SV=1 |
|
| 4.28e-217 | 1 | 522 | 3 | 511 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=mns1B PE=3 SV=1 |
|
| 1.01e-212 | 46 | 522 | 43 | 509 | Mannosyl-oligosaccharide alpha-1,2-mannosidase OS=Penicillium citrinum OX=5077 GN=MSDC PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
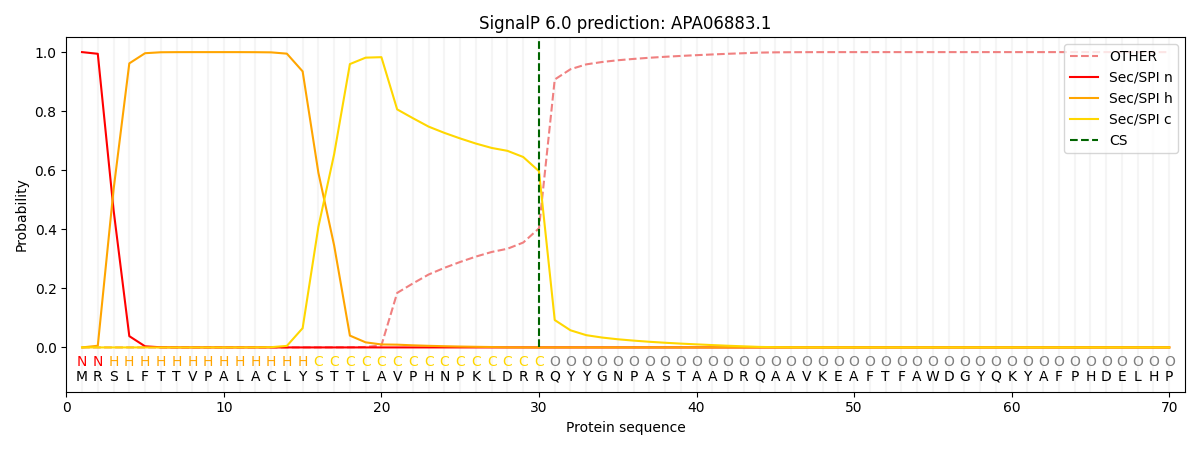
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000420 | 0.999544 | CS pos: 30-31. Pr: 0.5967 |
