You are browsing environment: FUNGIDB
CAZyme Information: AO090701000885-T-p1
You are here: Home > Sequence: AO090701000885-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus oryzae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus oryzae | |||||||||||
| CAZyme ID | AO090701000885-T-p1 | |||||||||||
| CAZy Family | GT62 | |||||||||||
| CAZyme Description | Alpha-L-arabinofuranosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.55:25 | 3.2.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH62 | 28 | 298 | 2e-135 | 0.9964028776978417 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 281639 | Glyco_hydro_62 | 0.0 | 28 | 298 | 1 | 272 | Glycosyl hydrolase family 62. Family of alpha -L-arabinofuranosidase (EC 3.2.1.55). This enzyme hydrolyzed aryl alpha-L-arabinofuranosides and cleaves arabinosyl side chains from arabinoxylan and arabinan. |
| 350101 | GH62 | 1.58e-166 | 30 | 326 | 1 | 304 | Glycosyl hydrolase family 62, characterized arabinofuranosidases. The glycosyl hydrolase family 62 (GH62) includes eukaryotic (mostly fungal) and prokaryotic enzymes which are characterized arabinofuranosidases (alpha-L-arabinofuranosidases; EC 3.2.1.55) that specifically cleave either alpha-1,2 or alpha-1,3-L-arabinofuranose side chains from xylans. These enzymes show significantly different substrate preference with rather low specific activity towards natural substrates and differ in catalytic efficiency. They do not act on xylose moieties in xylan that are adorned with an arabinose side chain at both O2 and O3 positions, nor do they display any non-specific arabinofuranosidase activity. The synergistic action in biomass degradation makes GH62 promising candidates for biotechnological improvements of biofuel production and in various biorefinery applications. These enzymes also contain carbohydrate binding modules (CBMs) that bind cellulose or xylan. |
| 350092 | GH_F | 4.45e-37 | 53 | 306 | 1 | 251 | Glycosyl hydrolase families 43 and 62 form CAZY clan GH-F. This glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) includes family 43 (GH43) and 62 (GH62). GH43 includes enzymes with beta-xylosidase (EC 3.2.1.37), beta-1,3-xylosidase (EC 3.2.1.-), alpha-L-arabinofuranosidase (EC 3.2.1.55), arabinanase (EC 3.2.1.99), xylanase (EC 3.2.1.8), endo-alpha-L-arabinanases (beta-xylanases) and galactan 1,3-beta-galactosidase (EC 3.2.1.145) activities. GH62 includes enzymes characterized as arabinofuranosidases (alpha-L-arabinofuranosidases; EC 3.2.1.55) that specifically cleave either alpha-1,2 or alpha-1,3-L-arabinofuranose side chains from xylans. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many of the enzymes in this family display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. GH62 are also predicted to be inverting enzymes. A common structural feature of both, GH43 and GH62 enzymes, is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| 350097 | GH43_Bt3655-like | 5.25e-05 | 142 | 267 | 125 | 243 | Glycosyl hydrolase family 43 protein such as Bacteroides thetaiotaomicron VPI-5482 arabinofuranosidase Bt3655. This glycosyl hydrolase family 43 (GH43)-like family includes the characterized arabinofuranosidases (EC 3.2.1.55): Bacteroides thetaiotaomicron VPI-5482 (Bt3655;BT_3655) and Penicillium chrysogenum 31B Abf43B, as well as Bifidobacterium adolescentis ATCC 15703 beta-xylosidase (EC 3.2.1.37) BAD_1527. It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 includes enzymes with beta-xylosidase (EC 3.2.1.37), beta-1,3-xylosidase (EC 3.2.1.-), alpha-L-arabinofuranosidase (EC 3.2.1.55), arabinanase (EC 3.2.1.99), xylanase (EC 3.2.1.8), endo-alpha-L-arabinanases (beta-xylanases) and galactan 1,3-beta-galactosidase (EC 3.2.1.145) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| 350115 | GH43_FsAxh1-like | 0.008 | 96 | 253 | 69 | 210 | Glycosyl hydrolase family 43 such as Fibrobacter succinogenes subsp. succinogenes S85 arabinoxylan alpha-L-arabinofuranosidase. This glycosyl hydrolase family 43 (GH43) includes mostly enzymes that have been annotated as having beta-1,4-xylosidase (beta-D-xylosidase; xylan 1,4-beta-xylosidase; EC 3.2.1.37) activity. They are part of an array of hemicellulases that are involved in the final breakdown of plant cell-wall whereby they degrade xylan. They hydrolyze beta-1,4 glycosidic bonds between two xylose units in short xylooligosaccharides. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. This subfamily includes the characterized Clostridium stercorarium F-9 beta-xylosidase Xyl43B. It also includes Humicola insolens AXHd3 (HiAXHd3), a GH43 arabinofuranosidase (EC 3.2.1.55) that hydrolyzes O3-linked arabinose of doubly substituted xylans, a feature of the polysaccharide that is recalcitrant to degradation. It possesses an additional C-terminal beta-sandwich domain such that the interface between the domains comprises a xylan binding cleft that houses the active site pocket. The HiAXHd3 active site is tuned to hydrolyze arabinofuranosyl or xylosyl linkages, and the topology of the distal regions of the substrate binding surface confers specificity. It also includes Fibrobacter succinogenes subsp. succinogenes S85 arabinoxylan alpha-L-arabinofuranosidase (Axh1;Fisuc_1769;FSU_2269), Paenibacillus sp. E18 alpha-L-arabinofuranosidase (Abf43A), Bifidobacterium adolescentis ATCC 15703 double substituted xylan alpha-1,3-L-specific arabinofuranosidase d3 (AXHd3;AXH-d3;BaAXH-d3;BAD_0301;E-AFAM2), and Chrysosporium lucknowense C1 arabinoxylan hydrolase / double substituted xylan alpha-1,3-L-arabinofuranosidase (Abn7;AXHd). A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.10e-244 | 1 | 328 | 1 | 328 | |
| 2.10e-244 | 1 | 328 | 1 | 328 | |
| 2.10e-244 | 1 | 328 | 1 | 328 | |
| 2.10e-244 | 1 | 328 | 1 | 328 | |
| 1.22e-243 | 1 | 328 | 1 | 328 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.93e-179 | 26 | 327 | 25 | 328 | Structure of an alpha-L-arabinofuranosidase (GH62) from Aspergillus nidulans [Aspergillus nidulans FGSC A4] |
|
| 1.44e-171 | 28 | 326 | 26 | 324 | Structure of a Talaromyces pinophilus GH62 Arabinofuranosidase in complex with AraDNJ at 1.25A resolution [Talaromyces pinophilus],6F1J_B Structure of a Talaromyces pinophilus GH62 Arabinofuranosidase in complex with AraDNJ at 1.25A resolution [Talaromyces pinophilus] |
|
| 7.82e-170 | 28 | 327 | 139 | 437 | Crystal Structure of Streptomyces coelicolor alpha-L-arabinofuranosidase [Streptomyces coelicolor A3(2)],3WMZ_A Crystal Structure of Streptomyces coelicolor alpha-L-arabinofuranosidase ethylmercury derivative [Streptomyces coelicolor A3(2)],3WN0_A Crystal Structure of Streptomyces coelicolor alpha-L-arabinofuranosidase in complex with L-arabinose [Streptomyces coelicolor A3(2)],3WN1_A Crystal Structure of Streptomyces coelicolor alpha-L-arabinofuranosidase in complex with xylotriose [Streptomyces coelicolor A3(2)],3WN2_A Crystal Structure of Streptomyces coelicolor alpha-L-arabinofuranosidase in complex with xylohexaose [Streptomyces coelicolor A3(2)] |
|
| 9.03e-169 | 28 | 327 | 20 | 318 | Structure of Fungal GH62 from Thielavia terretris [Thermothielavioides terrestris NRRL 8126] |
|
| 4.30e-166 | 23 | 327 | 78 | 383 | Crystal structure of SthAraf62A, a GH62 family alpha-L-arabinofuranosidase from Streptomyces thermoviolaceus, in the apoprotein form [Streptomyces thermoviolaceus],4O8O_A Crystal structure of SthAraf62A, a GH62 family alpha-L-arabinofuranosidase from Streptomyces thermoviolaceus, bound to alpha-L-arabinose [Streptomyces thermoviolaceus],4O8P_A Crystal structure of SthAraf62A, a GH62 family alpha-L-arabinofuranosidase from Streptomyces thermoviolaceus, bound to xylotetraose [Streptomyces thermoviolaceus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.73e-245 | 1 | 328 | 1 | 328 | Probable alpha-L-arabinofuranosidase axhA OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=axhA PE=3 SV=1 |
|
| 1.25e-243 | 1 | 328 | 1 | 328 | Alpha-L-arabinofuranosidase axhA OS=Aspergillus sojae OX=41058 GN=axhA PE=2 SV=1 |
|
| 4.86e-202 | 25 | 327 | 25 | 327 | Probable alpha-L-arabinofuranosidase axhA-1 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=axhA-1 PE=3 SV=2 |
|
| 1.43e-196 | 20 | 328 | 18 | 326 | Probable alpha-L-arabinofuranosidase axhA OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=axhA PE=3 SV=1 |
|
| 3.66e-178 | 26 | 327 | 22 | 325 | Alpha-L-arabinofuranosidase axhA-2 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=axhA-2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
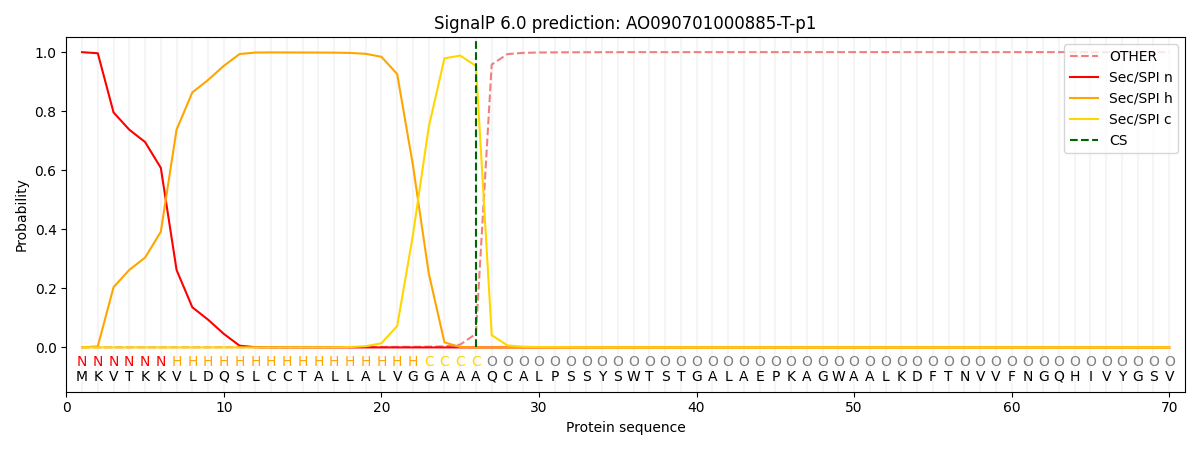
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000897 | 0.999057 | CS pos: 26-27. Pr: 0.9531 |
