You are browsing environment: FUNGIDB
CAZyme Information: AN3566-T-p1
You are here: Home > Sequence: AN3566-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
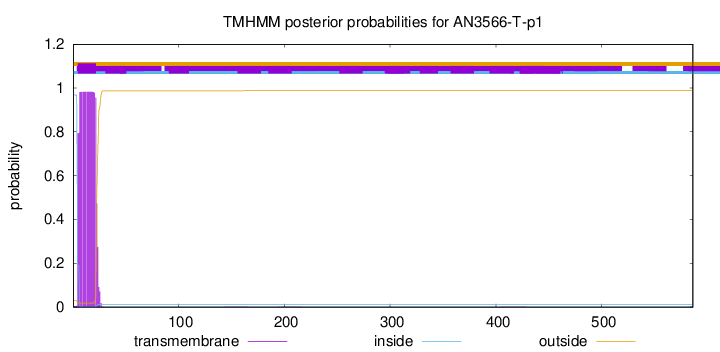
TMHMM annotations
Basic Information help
| Species | Aspergillus nidulans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus nidulans | |||||||||||
| CAZyme ID | AN3566-T-p1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | alpha-1,2-mannosidase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:7 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 117 | 582 | 8.1e-162 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 0.0 | 117 | 582 | 1 | 452 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.34e-134 | 108 | 582 | 70 | 517 | glycoside hydrolase family 47 protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 586 | 1 | 586 | |
| 0.0 | 1 | 586 | 1 | 586 | |
| 0.0 | 1 | 586 | 1 | 586 | |
| 0.0 | 1 | 586 | 1 | 586 | |
| 0.0 | 1 | 586 | 1 | 586 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.82e-70 | 108 | 584 | 35 | 509 | Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KKT_B Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KRE_A Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRE_B Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_A Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_B Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum] |
|
| 8.82e-70 | 108 | 584 | 3 | 474 | Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI8_B Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI9_A Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum],2RI9_B Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum] |
|
| 1.41e-68 | 110 | 581 | 5 | 449 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 1.64e-68 | 110 | 581 | 10 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 1.73e-68 | 110 | 581 | 10 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.79e-75 | 106 | 582 | 94 | 536 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS2 OS=Arabidopsis thaliana OX=3702 GN=MNS2 PE=1 SV=1 |
|
| 4.05e-75 | 108 | 582 | 38 | 506 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase ARB_00035 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_00035 PE=1 SV=1 |
|
| 1.09e-74 | 108 | 583 | 195 | 646 | Mannosyl-oligosaccharide alpha-1,2-mannosidase IA OS=Spodoptera frugiperda OX=7108 PE=1 SV=1 |
|
| 4.01e-73 | 106 | 582 | 93 | 535 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS1 OS=Arabidopsis thaliana OX=3702 GN=MNS1 PE=1 SV=1 |
|
| 9.24e-73 | 102 | 582 | 27 | 507 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=mns1B PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.989184 | 0.010827 |