You are browsing environment: FUNGIDB
CAZyme Information: ALNC14_100670:RNA-p1
You are here: Home > Sequence: ALNC14_100670:RNA-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
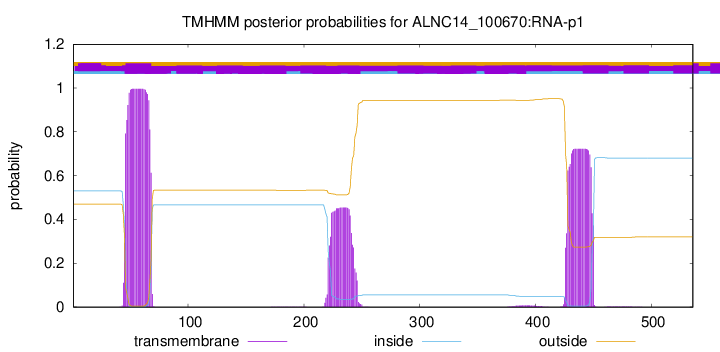
TMHMM annotations
Basic Information help
| Species | Albugo laibachii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Albuginaceae; Albugo; Albugo laibachii | |||||||||||
| CAZyme ID | ALNC14_100670:RNA-p1 | |||||||||||
| CAZy Family | GH81 | |||||||||||
| CAZyme Description | 5A endo1 putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 140 | 488 | 3.3e-140 | 0.997093023255814 |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 536 | 1 | 536 | |
| 2.16e-187 | 7 | 535 | 11 | 540 | |
| 1.57e-98 | 6 | 535 | 103 | 627 | |
| 1.57e-98 | 6 | 535 | 103 | 627 | |
| 8.79e-95 | 11 | 535 | 113 | 632 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.66e-19 | 137 | 526 | 55 | 410 | Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHO_B Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHO_C Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
|
| 8.30e-19 | 137 | 526 | 55 | 410 | Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHM_B Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHM_C Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
|
| 8.30e-19 | 137 | 526 | 55 | 410 | Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHN_B Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHN_C Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
|
| 1.11e-18 | 137 | 526 | 55 | 410 | Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_B Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_C Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3] |
|
| 1.44e-18 | 137 | 526 | 22 | 377 | The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_B The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_C The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3W6L_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999975 | 0.000066 |