You are browsing environment: FUNGIDB
CAZyme Information: AKAW_05190-t41_1-p1
You are here: Home > Sequence: AKAW_05190-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus luchuensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus luchuensis | |||||||||||
| CAZyme ID | AKAW_05190-t41_1-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | cutinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.1.1.74:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE5 | 47 | 223 | 1.6e-45 | 0.9894179894179894 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395860 | Cutinase | 1.03e-62 | 47 | 224 | 1 | 173 | Cutinase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.48e-167 | 1 | 257 | 1 | 257 | |
| 6.48e-167 | 1 | 257 | 1 | 257 | |
| 4.02e-153 | 1 | 257 | 1 | 262 | |
| 4.02e-153 | 1 | 257 | 1 | 262 | |
| 1.27e-110 | 1 | 227 | 1 | 224 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.71e-69 | 26 | 224 | 2 | 196 | Crystal structure of Aspergillus oryzae cutinase [Aspergillus oryzae] |
|
| 7.84e-68 | 41 | 224 | 5 | 187 | Structure of Aspergillus oryzae cutinase expressed in Pichia pastoris, crystallized in the presence of Paraoxon [Aspergillus oryzae] |
|
| 1.34e-65 | 37 | 224 | 3 | 193 | Crystal structure of a biodegradable plastic-degrading cutinase from Paraphoma sp. B47-9. [Paraphoma sp. B47-9],7CY3_B Crystal structure of a biodegradable plastic-degrading cutinase from Paraphoma sp. B47-9. [Paraphoma sp. B47-9],7CY9_A Crystal structure of a biodegradable plastic-degrading cutinase from Paraphoma sp. B47-9 solved by getting the phase from anomalous scattering of uncovalently coordinated arsenic (cacodylate). [Paraphoma sp. B47-9],7CY9_B Crystal structure of a biodegradable plastic-degrading cutinase from Paraphoma sp. B47-9 solved by getting the phase from anomalous scattering of uncovalently coordinated arsenic (cacodylate). [Paraphoma sp. B47-9] |
|
| 3.05e-65 | 26 | 224 | 2 | 198 | Chain A, cutinase [Malbranchea cinnamomea] |
|
| 4.22e-64 | 37 | 226 | 3 | 194 | Humicola insolens cutinase [Humicola insolens],4OYY_B Humicola insolens cutinase [Humicola insolens],4OYY_C Humicola insolens cutinase [Humicola insolens],4OYY_D Humicola insolens cutinase [Humicola insolens],4OYY_E Humicola insolens cutinase [Humicola insolens],4OYY_F Humicola insolens cutinase [Humicola insolens],4OYY_G Humicola insolens cutinase [Humicola insolens],4OYY_H Humicola insolens cutinase [Humicola insolens],4OYY_I Humicola insolens cutinase [Humicola insolens],4OYY_J Humicola insolens cutinase [Humicola insolens],4OYY_K Humicola insolens cutinase [Humicola insolens],4OYY_L Humicola insolens cutinase [Humicola insolens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.13e-154 | 1 | 257 | 1 | 262 | Probable cutinase 1 OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=An14g02170 PE=3 SV=1 |
|
| 2.26e-111 | 1 | 227 | 1 | 224 | Probable cutinase 3 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=AO090011000113 PE=3 SV=1 |
|
| 4.55e-111 | 1 | 227 | 1 | 224 | Probable cutinase 1 OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=AFLA_039350 PE=3 SV=1 |
|
| 1.71e-98 | 8 | 224 | 6 | 219 | Cutinase 3 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cut3 PE=2 SV=1 |
|
| 1.41e-96 | 8 | 225 | 6 | 216 | Probable cutinase 3 OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=NFIA_030250 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
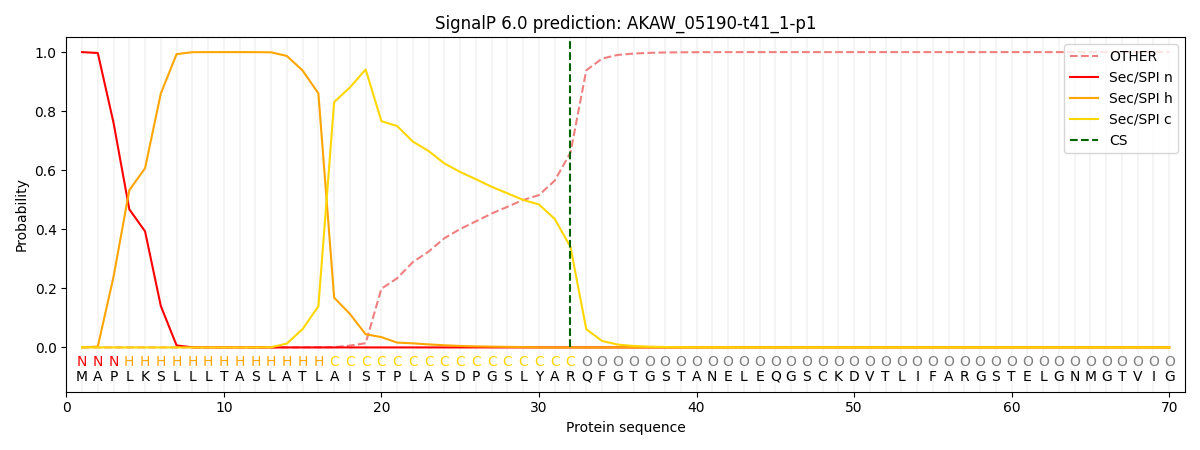
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000284 | 0.999698 | CS pos: 32-33. Pr: 0.3402 |
