You are browsing environment: FUNGIDB
CAZyme Information: AGR57_6095T0-p1
You are here: Home > Sequence: AGR57_6095T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phanerochaete chrysosporium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Phanerochaetaceae; Phanerochaete; Phanerochaete chrysosporium | |||||||||||
| CAZyme ID | AGR57_6095T0-p1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | FAD-binding domain-containing protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 28 | 457 | 3.1e-84 | 0.982532751091703 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223354 | GlcD | 4.11e-31 | 8 | 461 | 5 | 459 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 396238 | FAD_binding_4 | 3.59e-28 | 38 | 176 | 1 | 139 | FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidizes the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. |
| 215242 | PLN02441 | 1.58e-26 | 41 | 198 | 68 | 232 | cytokinin dehydrogenase |
| 223882 | MurB | 4.90e-09 | 38 | 206 | 21 | 182 | UDP-N-acetylenolpyruvoylglucosamine reductase [Cell wall/membrane/envelope biogenesis]. |
| 273751 | FAD_lactone_ox | 1.22e-07 | 44 | 209 | 21 | 183 | sugar 1,4-lactone oxidases. This model represents a family of at least two different sugar 1,4 lactone oxidases, both involved in synthesizing ascorbic acid or a derivative. These include L-gulonolactone oxidase (EC 1.1.3.8) from rat and D-arabinono-1,4-lactone oxidase (EC 1.1.3.37) from Saccharomyces cerevisiae. Members are proposed to have the cofactor FAD covalently bound at a site specified by Prosite motif PS00862; OX2_COVAL_FAD; 1. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.90e-29 | 41 | 454 | 66 | 484 | |
| 1.16e-24 | 41 | 211 | 23 | 193 | |
| 1.94e-24 | 9 | 455 | 26 | 463 | |
| 3.14e-23 | 41 | 237 | 167 | 365 | |
| 5.25e-22 | 41 | 237 | 65 | 262 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.27e-61 | 11 | 454 | 19 | 453 | The crystal structure of EncM V135T mutant [Streptomyces maritimus],6FYG_B The crystal structure of EncM V135T mutant [Streptomyces maritimus],6FYG_C The crystal structure of EncM V135T mutant [Streptomyces maritimus],6FYG_D The crystal structure of EncM V135T mutant [Streptomyces maritimus] |
|
| 6.81e-61 | 11 | 454 | 19 | 453 | Crystal Structure of EncM (crystallized with 4 mM NADPH) [Streptomyces maritimus],4XLO_B Crystal Structure of EncM (crystallized with 4 mM NADPH) [Streptomyces maritimus],4XLO_C Crystal Structure of EncM (crystallized with 4 mM NADPH) [Streptomyces maritimus],4XLO_D Crystal Structure of EncM (crystallized with 4 mM NADPH) [Streptomyces maritimus],6FOQ_A The crystal structure of EncM complexed with dioxygen under 15 bar of oxygen pressure. [Streptomyces maritimus],6FOQ_B The crystal structure of EncM complexed with dioxygen under 15 bar of oxygen pressure. [Streptomyces maritimus],6FOQ_C The crystal structure of EncM complexed with dioxygen under 15 bar of oxygen pressure. [Streptomyces maritimus],6FOQ_D The crystal structure of EncM complexed with dioxygen under 15 bar of oxygen pressure. [Streptomyces maritimus],6FOW_A The crystal structure of EncM complexed with dioxygen under 10 bar of oxygen pressure. [Streptomyces maritimus],6FOW_B The crystal structure of EncM complexed with dioxygen under 10 bar of oxygen pressure. [Streptomyces maritimus],6FOW_C The crystal structure of EncM complexed with dioxygen under 10 bar of oxygen pressure. [Streptomyces maritimus],6FOW_D The crystal structure of EncM complexed with dioxygen under 10 bar of oxygen pressure. [Streptomyces maritimus],6FP3_A The crystal structure of EncM complexed with dioxygen under 5 bar of oxygen pressure. [Streptomyces maritimus],6FP3_B The crystal structure of EncM complexed with dioxygen under 5 bar of oxygen pressure. [Streptomyces maritimus],6FP3_C The crystal structure of EncM complexed with dioxygen under 5 bar of oxygen pressure. [Streptomyces maritimus],6FP3_D The crystal structure of EncM complexed with dioxygen under 5 bar of oxygen pressure. [Streptomyces maritimus],6FY8_A The crystal structure of EncM bromide soak [Streptomyces maritimus],6FY9_A The crystal structure of EncM complex with xenon under 15 bars Xe pressure [Streptomyces maritimus],6FYA_A The crystal structure of EncM under anaerobic conditions [Streptomyces maritimus],6FYA_B The crystal structure of EncM under anaerobic conditions [Streptomyces maritimus] |
|
| 7.47e-61 | 11 | 454 | 23 | 457 | The crystal structure of EncM [Streptomyces maritimus],3W8W_B The crystal structure of EncM [Streptomyces maritimus],3W8X_A The complex structure of EncM with trifluorotriketide [Streptomyces maritimus],3W8X_B The complex structure of EncM with trifluorotriketide [Streptomyces maritimus],3W8Z_A The complex structure of EncM with hydroxytetraketide [Streptomyces maritimus],3W8Z_B The complex structure of EncM with hydroxytetraketide [Streptomyces maritimus] |
|
| 9.53e-61 | 11 | 454 | 19 | 453 | The crystal structure of EncM V135M mutant [Streptomyces maritimus],6FYF_B The crystal structure of EncM V135M mutant [Streptomyces maritimus],6FYF_C The crystal structure of EncM V135M mutant [Streptomyces maritimus],6FYF_D The crystal structure of EncM V135M mutant [Streptomyces maritimus] |
|
| 1.33e-60 | 11 | 454 | 19 | 453 | The crystal structure of EncM L144M mutant [Streptomyces maritimus],6FYB_B The crystal structure of EncM L144M mutant [Streptomyces maritimus],6FYB_C The crystal structure of EncM L144M mutant [Streptomyces maritimus],6FYB_D The crystal structure of EncM L144M mutant [Streptomyces maritimus],6FYC_A The crystal structure of EncM L144M mutant complex with dioxygen under 15 bars O2 pressure [Streptomyces maritimus],6FYC_B The crystal structure of EncM L144M mutant complex with dioxygen under 15 bars O2 pressure [Streptomyces maritimus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.00e-120 | 14 | 462 | 15 | 451 | FAD-linked oxidoreductase pyvE OS=Aspergillus violaceofuscus (strain CBS 115571) OX=1450538 GN=pyvE PE=3 SV=1 |
|
| 5.28e-43 | 23 | 454 | 32 | 441 | FAD-linked oxidoreductase DDB_G0289697 OS=Dictyostelium discoideum OX=44689 GN=DDB_G0289697 PE=2 SV=1 |
|
| 3.43e-41 | 23 | 454 | 63 | 496 | FAD-linked oxidoreductase OXR1 OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=OXR1 PE=1 SV=1 |
|
| 5.60e-41 | 13 | 464 | 14 | 457 | (R)-6-hydroxynicotine oxidase OS=Paenarthrobacter nicotinovorans OX=29320 GN=6-hdno PE=1 SV=2 |
|
| 9.98e-39 | 29 | 454 | 65 | 489 | FAD-linked oxidoreductase srdI OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=srdI PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
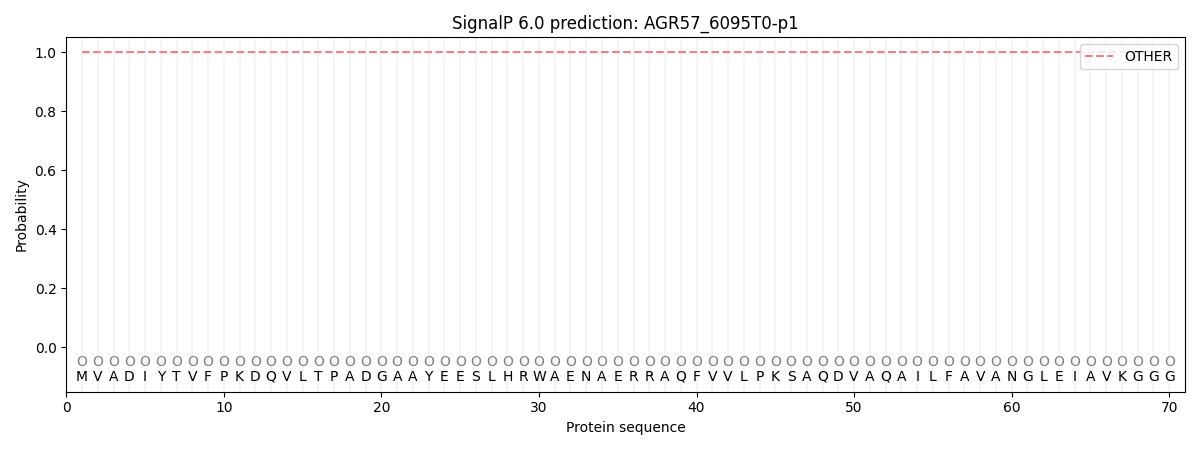
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000040 | 0.000001 |
