You are browsing environment: FUNGIDB
CAZyme Information: AGR57_4581T0-p1
You are here: Home > Sequence: AGR57_4581T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phanerochaete chrysosporium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Phanerochaetaceae; Phanerochaete; Phanerochaete chrysosporium | |||||||||||
| CAZyme ID | AGR57_4581T0-p1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Carbohydrate-Binding Module Family 1 / Carbohydrate Esterase Family 16 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE16 | 98 | 354 | 2e-81 | 0.9887640449438202 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 238882 | fatty_acyltransferase_like | 1.42e-58 | 100 | 355 | 5 | 268 | Fatty acyltransferase-like subfamily of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. Might catalyze fatty acid transfer between phosphatidylcholine and sterols. |
| 238875 | SGNH_plant_lipase_like | 4.51e-25 | 98 | 354 | 4 | 311 | SGNH_plant_lipase_like, a plant specific subfamily of the SGNH-family of hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| 238883 | Triacylglycerol_lipase_like | 4.69e-12 | 98 | 358 | 5 | 281 | Triacylglycerol lipase-like subfamily of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. Members of this subfamily might hydrolyze triacylglycerol into diacylglycerol and fatty acid anions. |
| 395595 | CBM_1 | 4.47e-11 | 23 | 60 | 2 | 29 | Fungal cellulose binding domain. |
| 225780 | COG3240 | 2.33e-10 | 76 | 329 | 12 | 284 | Phospholipase/lecithinase/hemolysin [Lipid transport and metabolism, General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.22e-131 | 17 | 361 | 21 | 373 | |
| 1.31e-128 | 12 | 361 | 16 | 373 | |
| 1.31e-128 | 12 | 361 | 16 | 373 | |
| 1.25e-125 | 8 | 361 | 11 | 361 | |
| 6.91e-107 | 16 | 361 | 28 | 383 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.36e-08 | 20 | 64 | 4 | 38 | Chain A, ENDOGLUCANASE EG-1 [Trichoderma reesei] |
|
| 1.04e-07 | 18 | 67 | 21 | 70 | Crystal structure of Endo-1,4-beta-xylanase-like protein from Acremonium chrysogenum [Acremonium chrysogenum ATCC 11550],5MRJ_B Crystal structure of Endo-1,4-beta-xylanase-like protein from Acremonium chrysogenum [Acremonium chrysogenum ATCC 11550] |
|
| 1.84e-06 | 20 | 64 | 2 | 36 | Chain A, C-TERMINAL DOMAIN OF CELLOBIOHYDROLASE I [Trichoderma reesei],2CBH_A Chain A, C-TERMINAL DOMAIN OF CELLOBIOHYDROLASE I [Trichoderma reesei],2MWJ_A Chain A, Exoglucanase 1 [Trichoderma reesei],2MWK_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X34_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X35_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X36_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X37_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X38_A Chain A, Exoglucanase 1 [Trichoderma reesei] |
|
| 2.72e-06 | 18 | 66 | 3 | 41 | Crystal Structure of Glucuronoyl Esterase from Cerrena unicolor covalent complex with the aldouronic acid UXXR [Cerrena unicolor],6RV8_B Crystal Structure of Glucuronoyl Esterase from Cerrena unicolor covalent complex with the aldouronic acid UXXR [Cerrena unicolor] |
|
| 7.60e-06 | 22 | 64 | 374 | 406 | GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8M_B GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8N_A GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872],6Q8N_B GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.26e-78 | 92 | 355 | 29 | 294 | Acetylesterase OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=aes1 PE=3 SV=1 |
|
| 4.02e-77 | 92 | 355 | 29 | 294 | Acetylesterase OS=Thermothelomyces thermophilus OX=78579 GN=aes1 PE=1 SV=1 |
|
| 6.37e-12 | 18 | 70 | 20 | 62 | 4-O-methyl-glucuronoyl methylesterase 1 OS=Phanerochaete chrysosporium (strain RP-78 / ATCC MYA-4764 / FGSC 9002) OX=273507 GN=e_gw1.18.61.1 PE=1 SV=1 |
|
| 2.85e-11 | 11 | 83 | 10 | 72 | Manganese dependent endoglucanase Eg5A OS=Phanerodontia chrysosporium OX=2822231 GN=Eg5A PE=1 SV=2 |
|
| 8.51e-11 | 4 | 73 | 4 | 63 | Probable 1,4-beta-D-glucan cellobiohydrolase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=cbhC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
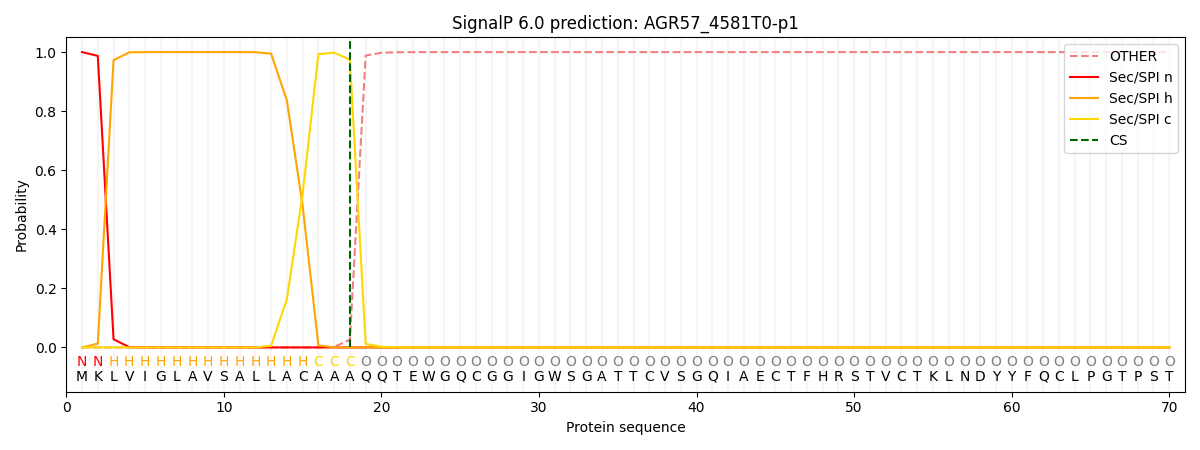
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000196 | 0.999777 | CS pos: 18-19. Pr: 0.9742 |
