You are browsing environment: FUNGIDB
CAZyme Information: AGR57_1889T0-p1
You are here: Home > Sequence: AGR57_1889T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phanerochaete chrysosporium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Phanerochaetaceae; Phanerochaete; Phanerochaete chrysosporium | |||||||||||
| CAZyme ID | AGR57_1889T0-p1 | |||||||||||
| CAZy Family | GH152 | |||||||||||
| CAZyme Description | Glycosyltransferase Family 2 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 594 | 716 | 2e-17 | 0.7176470588235294 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 178298 | PLN02695 | 7.97e-96 | 2 | 306 | 17 | 335 | GDP-D-mannose-3',5'-epimerase |
| 187581 | GME-like_SDR_e | 1.40e-92 | 8 | 306 | 3 | 318 | Arabidopsis thaliana GDP-mannose-3',5'-epimerase (GME)-like, extended (e) SDRs. This subgroup of NDP-sugar epimerase/dehydratases are extended SDRs; they have the characteristic active site tetrad, and an NAD-binding motif: TGXXGXX[AG], which is a close match to the canonical NAD-binding motif. Members include Arabidopsis thaliana GDP-mannose-3',5'-epimerase (GME) which catalyzes the epimerization of two positions of GDP-alpha-D-mannose to form GDP-beta-L-galactose. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
| 223528 | WcaG | 2.36e-61 | 8 | 300 | 3 | 308 | Nucleoside-diphosphate-sugar epimerase [Cell wall/membrane/envelope biogenesis]. |
| 187566 | UDP_AE_SDR_e | 5.02e-54 | 8 | 300 | 2 | 304 | UDP-N-acetylglucosamine 4-epimerase, extended (e) SDRs. This subgroup contains UDP-N-acetylglucosamine 4-epimerase of Pseudomonas aeruginosa, WbpP, an extended SDR, that catalyzes the NAD+ dependent conversion of UDP-GlcNAc and UDPGalNA to UDP-Glc and UDP-Gal. This subgroup has the characteristic active site tetrad and NAD-binding motif of the extended SDRs. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
| 187541 | UGD_SDR_e | 5.57e-45 | 8 | 299 | 3 | 303 | UDP-glucuronate decarboxylase (UGD) and related proteins, extended (e) SDRs. UGD catalyzes the formation of UDP-xylose from UDP-glucuronate; it is an extended-SDR, and has the characteristic glycine-rich NAD-binding pattern, TGXXGXXG, and active site tetrad. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 7.06e-316 | 7 | 930 | 18 | 1007 | |
| 5.47e-310 | 7 | 930 | 18 | 1007 | |
| 1.36e-295 | 7 | 930 | 19 | 1011 | |
| 1.50e-270 | 2 | 901 | 8 | 924 | |
| 7.82e-25 | 592 | 847 | 4 | 259 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.83e-64 | 2 | 306 | 26 | 344 | gdp-mannose-3', 5' -epimerase (arabidopsis thaliana), with gdp-alpha-d-mannose and gdp-beta-l-galactose bound in the active site. [Arabidopsis thaliana],2C59_B gdp-mannose-3', 5' -epimerase (arabidopsis thaliana), with gdp-alpha-d-mannose and gdp-beta-l-galactose bound in the active site. [Arabidopsis thaliana] |
|
| 1.78e-63 | 2 | 306 | 26 | 344 | gdp-mannose-3', 5' -epimerase (arabidopsis thaliana),k178r, with gdp-beta-l-gulose and gdp-4-keto-beta-l-gulose bound in active site. [Arabidopsis thaliana],2C54_B gdp-mannose-3', 5' -epimerase (arabidopsis thaliana),k178r, with gdp-beta-l-gulose and gdp-4-keto-beta-l-gulose bound in active site. [Arabidopsis thaliana] |
|
| 2.45e-63 | 2 | 306 | 26 | 344 | GDP-mannose-3', 5' -epimerase (Arabidopsis thaliana),Y174F, with GDP-beta-L-galactose bound in the active site [Arabidopsis thaliana],2C5A_B GDP-mannose-3', 5' -epimerase (Arabidopsis thaliana),Y174F, with GDP-beta-L-galactose bound in the active site [Arabidopsis thaliana] |
|
| 4.65e-63 | 2 | 306 | 26 | 344 | gdp-mannose-3', 5' -epimerase (arabidopsis thaliana), k217a, with gdp-alpha-d-mannose bound in the active site. [Arabidopsis thaliana],2C5E_B gdp-mannose-3', 5' -epimerase (arabidopsis thaliana), k217a, with gdp-alpha-d-mannose bound in the active site. [Arabidopsis thaliana] |
|
| 2.48e-28 | 8 | 304 | 3 | 310 | Crystal Structure of UDP-galactose 4-epimerase [Pyrobaculum calidifontis JCM 11548],3KO8_A Crystal Structure of UDP-galactose 4-epimerase [Pyrobaculum calidifontis JCM 11548] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.03e-65 | 2 | 306 | 26 | 343 | GDP-mannose 3,5-epimerase 1 OS=Oryza sativa subsp. japonica OX=39947 GN=GME-1 PE=1 SV=1 |
|
| 4.36e-65 | 2 | 306 | 19 | 336 | GDP-mannose 3,5-epimerase 2 OS=Oryza sativa subsp. japonica OX=39947 GN=GME-2 PE=2 SV=2 |
|
| 1.39e-64 | 2 | 306 | 26 | 343 | GDP-mannose 3,5-epimerase 1 OS=Oryza sativa subsp. indica OX=39946 GN=OsI_032456 PE=2 SV=1 |
|
| 3.33e-63 | 2 | 306 | 24 | 342 | GDP-mannose 3,5-epimerase OS=Arabidopsis thaliana OX=3702 GN=At5g28840 PE=1 SV=1 |
|
| 1.09e-25 | 1 | 299 | 12 | 315 | GDP-L-fucose synthase 1 OS=Arabidopsis thaliana OX=3702 GN=GER1 PE=1 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
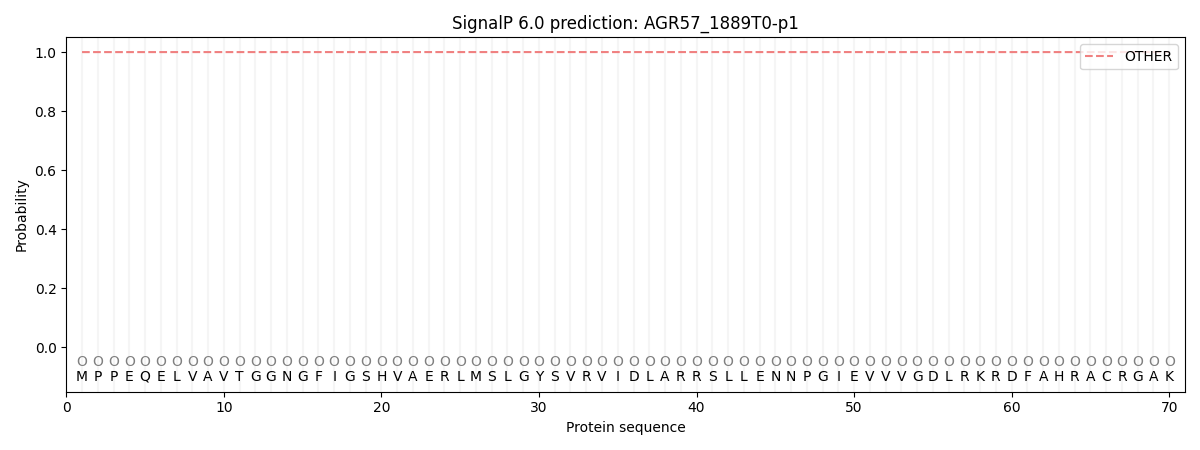
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000062 | 0.000000 |
