You are browsing environment: FUNGIDB
CAZyme Information: AGR57_10532T0-p1
You are here: Home > Sequence: AGR57_10532T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phanerochaete chrysosporium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Phanerochaetaceae; Phanerochaete; Phanerochaete chrysosporium | |||||||||||
| CAZyme ID | AGR57_10532T0-p1 | |||||||||||
| CAZy Family | AA2 | |||||||||||
| CAZyme Description | Glycoside Hydrolase Family 72 / Carbohydrate-Binding Module Family 43 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH72 | 281 | 600 | 3e-96 | 0.9839743589743589 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397351 | Glyco_hydro_72 | 2.73e-99 | 274 | 602 | 1 | 315 | Glucanosyltransferase. This is a family of glycosylphosphatidylinositol-anchored beta(1-3)glucanosyltransferases. The active site residues in the Aspergillus fumigatus example are the two glutamate residues at 160 and 261. |
| 400371 | X8 | 5.56e-19 | 647 | 729 | 1 | 76 | X8 domain. The X8 domain domain contains at least 6 conserved cysteine residues that presumably form three disulphide bridges. The domain is found in an Olive pollen allergen as well as at the C-terminus of several families of glycosyl hydrolases. This domain may be involved in carbohydrate binding. This domain is characteristic of GPI-anchored domains. |
| 399580 | SUR7 | 7.08e-14 | 10 | 248 | 5 | 200 | SUR7/PalI family. This family consists of several fungal-specific SUR7 proteins. Its activity regulates expression of RVS161, a homolog of human endophilin, suggesting a function for both in endocytosis. The protein carries four transmembrane domains and is thus likely to act as an anchoring protein for the eisosome to the plasma membrane. Eisosomes are the immobile protein complexes, that include the proteins Pil1 and Lsp1, which co-localize with sites of protein and lipid endocytosis at the plasma membrane. SUR7 protein may play a role in sporulation. This family also includes PalI which is part of a pH signal transduction cascade. Based on the similarity of PalI to the yeast Rim9 meiotic signal transduction component it has been suggested that PalI might be a membrane sensor for ambient pH. |
| 197867 | X8 | 4.94e-11 | 648 | 732 | 2 | 75 | Possibly involved in carbohydrate binding. The X8 domain, which may be involved in carbohydrate binding, is found in an Olive pollen antigen as well as at the C terminus of family 17 glycosyl hydrolases. It contains 6 conserved cysteine residues which presumably form three disulfide bridges. |
| 227605 | YqbO | 0.002 | 107 | 283 | 370 | 563 | Phage-related minor tail protein [Mobilome: prophages, transposons]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.91e-237 | 263 | 815 | 7 | 555 | |
| 8.28e-195 | 267 | 749 | 103 | 582 | |
| 1.19e-192 | 285 | 761 | 156 | 630 | |
| 2.61e-192 | 285 | 749 | 156 | 617 | |
| 9.03e-188 | 271 | 761 | 9 | 496 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.82e-71 | 290 | 722 | 42 | 466 | Saccharomyces cerevisiae Gas2p in complex with laminaripentaose [Saccharomyces cerevisiae],2W63_A Saccharomyces Cerevisiae Gas2p In Complex With Laminaritriose And Laminaritetraose [Saccharomyces cerevisiae],5O9O_A Crystal structure of ScGas2 in complex with compound 7. [Saccharomyces cerevisiae S288C],5O9P_A Crystal structure of Gas2 in complex with compound 10 [Saccharomyces cerevisiae S288C],5O9Q_A Crystal structure of ScGas2 in complex with compound 6 [Saccharomyces cerevisiae S288C],5O9R_A Crystal structure of ScGas2 in complex with compound 9 [Saccharomyces cerevisiae S288C],5O9Y_A Crystal structure of ScGas2 in complex with compound 11 [Saccharomyces cerevisiae S288C],5OA2_A Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_B Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_C Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA6_A Crystal structure of ScGas2 in complex with compound 12 [Saccharomyces cerevisiae S288C] |
|
| 1.28e-70 | 290 | 722 | 42 | 466 | Saccharomyces cerevisiae Gas2p apostructure (E176Q mutant) [Saccharomyces cerevisiae] |
|
| 1.28e-70 | 290 | 722 | 42 | 466 | SACCHAROMYCES CEREVISIAE GAS2P (E176Q MUTANT) IN COMPLEX WITH LAMINARITETRAOSE AND LAMINARIPENTAOSE [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.45e-100 | 290 | 728 | 45 | 467 | 1,3-beta-glucanosyltransferase gel4 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=gel4 PE=1 SV=1 |
|
| 1.45e-100 | 290 | 728 | 45 | 467 | 1,3-beta-glucanosyltransferase gel4 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gel4 PE=3 SV=1 |
|
| 6.98e-100 | 275 | 732 | 5 | 453 | 1,3-beta-glucanosyltransferase gel3 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gel3 PE=3 SV=1 |
|
| 7.59e-100 | 275 | 732 | 5 | 453 | 1,3-beta-glucanosyltransferase gel3 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=gel3 PE=1 SV=1 |
|
| 3.08e-98 | 290 | 730 | 34 | 453 | 1,3-beta-glucanosyltransferase GAS1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=GAS1 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
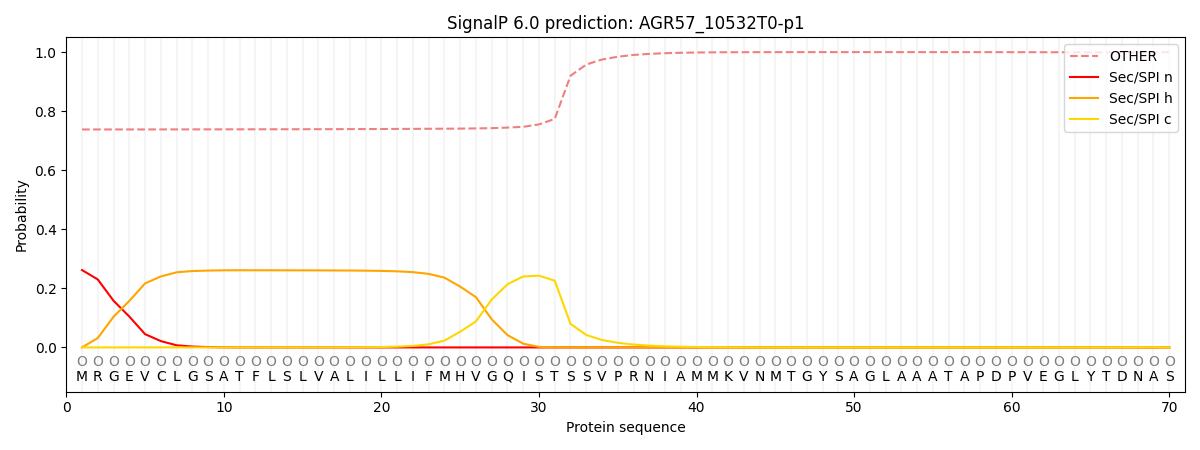
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.748006 | 0.252008 |
TMHMM Annotations download full data without filtering help
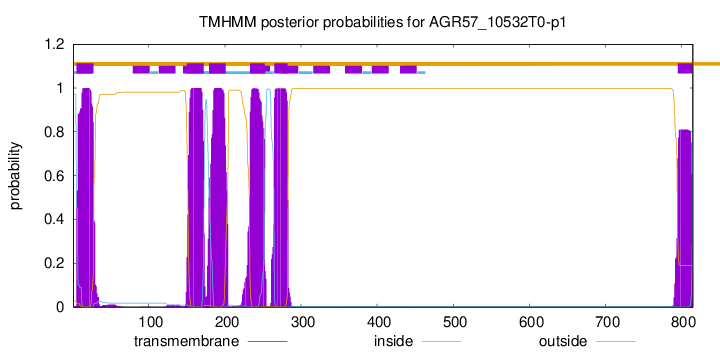
| Start | End |
|---|---|
| 5 | 27 |
| 150 | 172 |
| 179 | 201 |
| 233 | 252 |
| 265 | 282 |
| 795 | 814 |
