You are browsing environment: FUNGIDB
CAZyme Information: ACLA_074380-t26_1-p1
You are here: Home > Sequence: ACLA_074380-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus clavatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus clavatus | |||||||||||
| CAZyme ID | ACLA_074380-t26_1-p1 | |||||||||||
| CAZy Family | GH76 | |||||||||||
| CAZyme Description | conserved hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH131 | 18 | 256 | 5.4e-96 | 0.9921568627450981 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 408085 | GH131_N | 9.91e-89 | 18 | 251 | 1 | 251 | Glycoside hydrolase 131 catalytic N-terminal domain. This is the N-terminal domain found in glycoside hydrolase family 131 (GH131A) protein observed in Coprinopsis cinerea. GH131A exhibits bifunctional exo-beta-1,3-/-1,6- and endo-beta-1,4 activity toward beta-glucan. This domain is catalytic in nature though the catalytic mechanism of C. cinerea GH131A is different from that of typical glycosidases that use a pair of carboxylic acid residues as the catalytic residues. In the case of GH131A, Glu98 and His218 may form a catalytic dyad and Glu98 may activate His218 during catalysis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.92e-126 | 4 | 257 | 1 | 248 | |
| 4.92e-126 | 4 | 257 | 1 | 248 | |
| 4.92e-126 | 4 | 257 | 1 | 248 | |
| 4.92e-126 | 4 | 257 | 1 | 248 | |
| 1.29e-123 | 4 | 257 | 1 | 258 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.21e-99 | 18 | 257 | 3 | 243 | Chain A, Beta-glucanase [Podospora anserina],4LE3_B Chain B, Beta-glucanase [Podospora anserina],4LE3_C Chain C, Beta-glucanase [Podospora anserina],4LE3_D Chain D, Beta-glucanase [Podospora anserina],4LE4_A Chain A, Beta-glucanase [Podospora anserina],4LE4_B Chain B, Beta-glucanase [Podospora anserina],4LE4_C Chain C, Beta-glucanase [Podospora anserina],4LE4_D Chain D, Beta-glucanase [Podospora anserina] |
|
| 1.21e-95 | 17 | 252 | 2 | 229 | Chain A, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_B Chain B, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_C Chain C, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_D Chain D, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
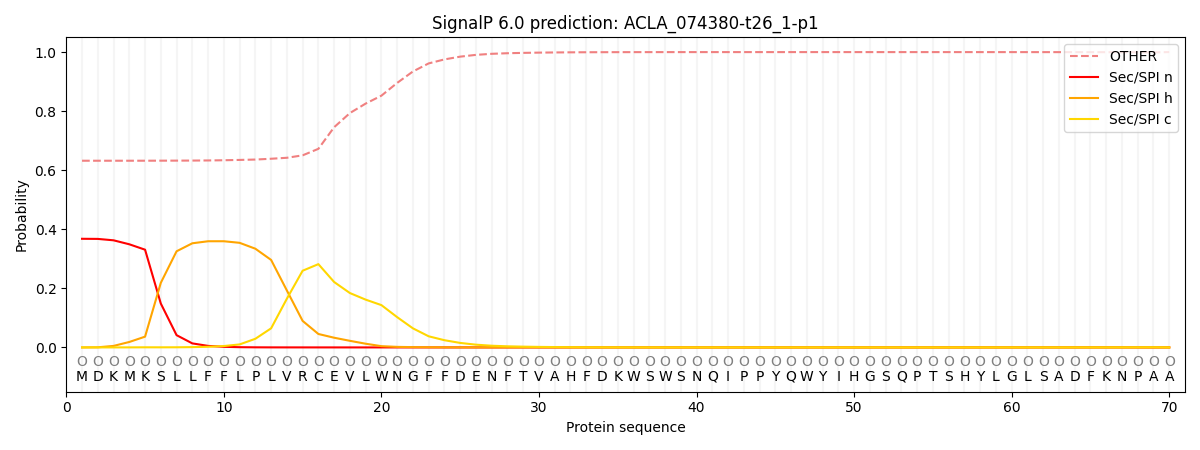
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.642057 | 0.357960 |
