You are browsing environment: FUNGIDB
CAZyme Information: ACLA_052760-t26_1-p1
You are here: Home > Sequence: ACLA_052760-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus clavatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus clavatus | |||||||||||
| CAZyme ID | ACLA_052760-t26_1-p1 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | glycosyl hydrolase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 88 | 312 | 9.9e-47 | 0.9675925925925926 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395747 | Glyco_hydro_3 | 4.42e-60 | 44 | 352 | 19 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| 224389 | BglX | 2.56e-54 | 56 | 350 | 29 | 309 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 235417 | PRK05337 | 2.45e-39 | 57 | 312 | 26 | 278 | beta-hexosaminidase; Provisional |
| 185053 | PRK15098 | 8.74e-11 | 44 | 356 | 64 | 354 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.17e-137 | 32 | 354 | 50 | 372 | |
| 3.42e-122 | 29 | 355 | 29 | 356 | |
| 3.45e-118 | 1 | 355 | 1 | 348 | |
| 1.16e-117 | 24 | 355 | 21 | 354 | |
| 5.53e-113 | 31 | 357 | 27 | 356 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.03e-34 | 31 | 357 | 17 | 341 | Structure of a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
|
| 5.70e-22 | 29 | 310 | 19 | 291 | Crystal structure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G1M_B Crystal structure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G2M_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G2M_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G3R_A Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G3R_B Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G5K_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5G5K_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
|
| 1.46e-21 | 29 | 310 | 19 | 291 | Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G5U_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_A Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_B Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5LY7_A Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5LY7_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
|
| 5.98e-21 | 47 | 355 | 76 | 395 | Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
|
| 2.55e-20 | 47 | 355 | 50 | 369 | Chain A, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.25e-104 | 34 | 355 | 28 | 350 | Uncharacterized secreted glycosyl hydrolase ARB_06359 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_06359 PE=1 SV=1 |
|
| 2.22e-24 | 35 | 313 | 4 | 278 | Beta-hexosaminidase OS=Psychromonas ingrahamii (strain 37) OX=357804 GN=nagZ PE=3 SV=1 |
|
| 9.01e-22 | 41 | 328 | 10 | 290 | Beta-hexosaminidase OS=Tolumonas auensis (strain DSM 9187 / TA4) OX=595494 GN=nagZ PE=3 SV=1 |
|
| 1.49e-21 | 52 | 312 | 23 | 281 | Beta-hexosaminidase OS=Nitrosomonas eutropha (strain DSM 101675 / C91 / Nm57) OX=335283 GN=nagZ PE=3 SV=1 |
|
| 1.46e-20 | 51 | 312 | 20 | 275 | Beta-hexosaminidase OS=Idiomarina loihiensis (strain ATCC BAA-735 / DSM 15497 / L2-TR) OX=283942 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
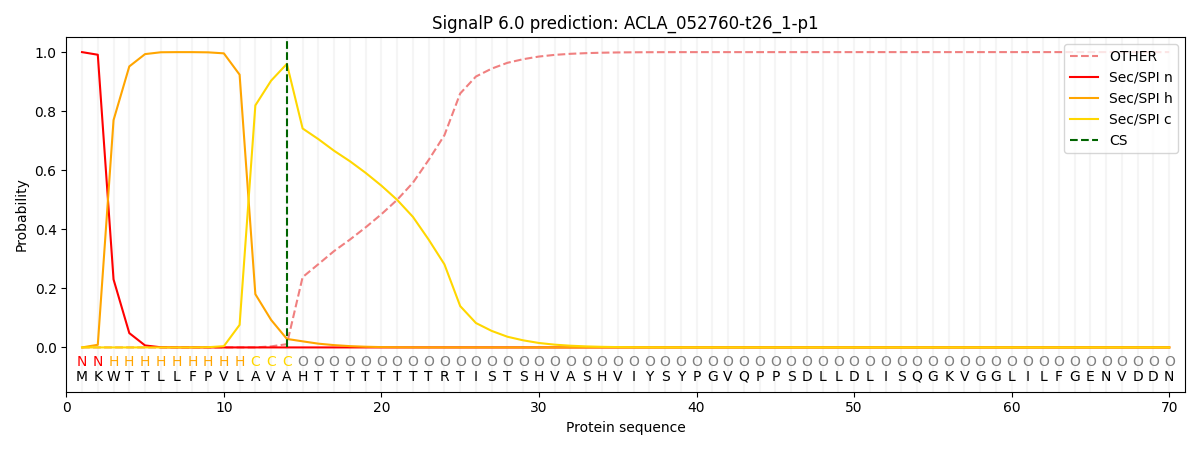
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000380 | 0.999582 | CS pos: 14-15. Pr: 0.9601 |
