You are browsing environment: FUNGIDB
CAZyme Information: ACLA_016840-t26_1-p1
You are here: Home > Sequence: ACLA_016840-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus clavatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus clavatus | |||||||||||
| CAZyme ID | ACLA_016840-t26_1-p1 | |||||||||||
| CAZy Family | AA8|AA3 | |||||||||||
| CAZyme Description | fungal chitosanase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.132:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH75 | 48 | 244 | 8.1e-70 | 0.990909090909091 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 399960 | Glyco_hydro_75 | 7.07e-56 | 100 | 244 | 1 | 165 | Fungal chitosanase of glycosyl hydrolase group 75. This family consists of several fungal chitosanase proteins. Chitin, xylan, 6-O-sulphated chitosan and O-carboxymethyl chitin are indigestible by chitosanase. EC:3.2.1.132. The mechanism is likely to be inverting, and the probable catalytic neutrophile base is Asp, with the probable catalytic proton donor being Glu. (see the Chitosanase web-page from CAZY). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.94e-130 | 8 | 319 | 6 | 311 | |
| 3.59e-122 | 21 | 316 | 20 | 304 | |
| 4.28e-121 | 11 | 313 | 10 | 302 | |
| 1.14e-119 | 1 | 313 | 2 | 296 | |
| 3.17e-115 | 4 | 316 | 4 | 307 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.53e-115 | 4 | 316 | 4 | 307 | Endo-chitosanase C OS=Aspergillus oryzae OX=5062 GN=csnC PE=1 SV=2 |
|
| 2.42e-110 | 4 | 316 | 4 | 316 | Endo-chitosanase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=csnC PE=3 SV=1 |
|
| 5.26e-22 | 100 | 242 | 78 | 235 | Endo-chitosanase B OS=Aspergillus oryzae OX=5062 GN=csnB PE=3 SV=1 |
|
| 5.04e-21 | 100 | 242 | 78 | 236 | Endo-chitosanase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=csnB PE=3 SV=1 |
|
| 5.45e-21 | 98 | 242 | 77 | 240 | Endo-chitosanase OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=csn PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
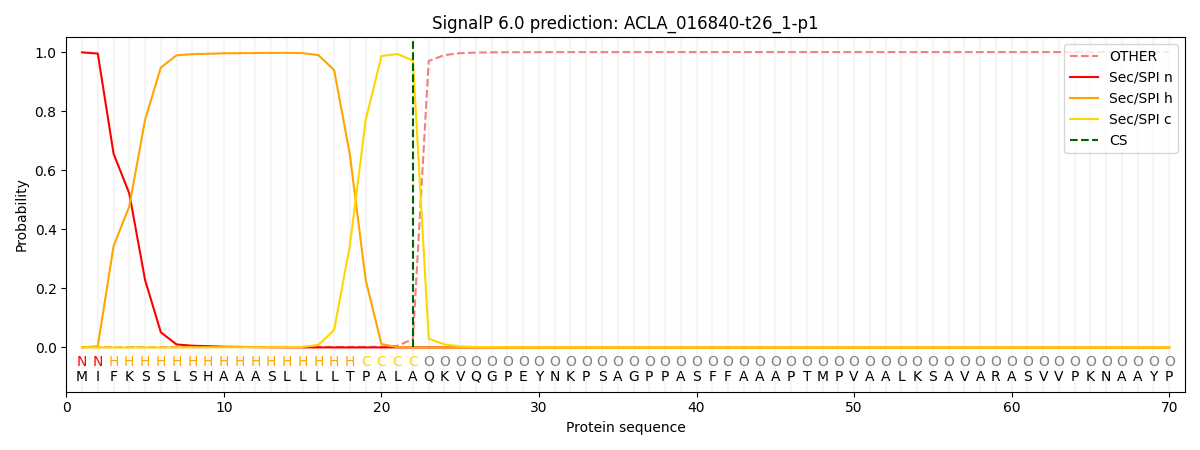
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.001890 | 0.998098 | CS pos: 22-23. Pr: 0.9716 |
