You are browsing environment: FUNGIDB
CAZyme Information: ACLA_002970-t26_1-p1
You are here: Home > Sequence: ACLA_002970-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
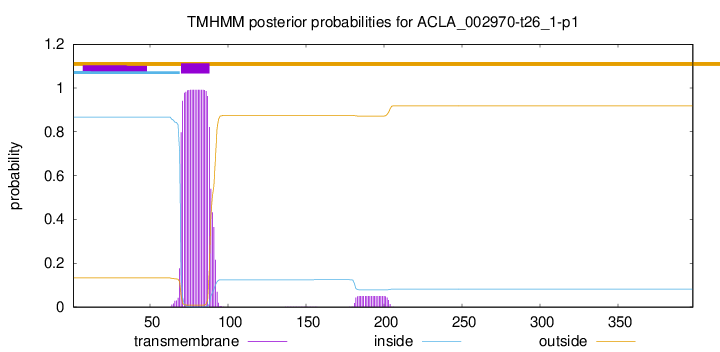
TMHMM annotations
Basic Information help
| Species | Aspergillus clavatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus clavatus | |||||||||||
| CAZyme ID | ACLA_002970-t26_1-p1 | |||||||||||
| CAZy Family | AA2 | |||||||||||
| CAZyme Description | endo-1,3(4)-beta-glucanase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 144 | 373 | 1.5e-84 | 0.982532751091703 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185690 | GH16_fungal_Lam16A_glucanase | 9.40e-137 | 104 | 394 | 2 | 293 | fungal 1,3(4)-beta-D-glucanases, similar to Phanerochaete chrysosporium laminarinase 16A. Group of fungal 1,3(4)-beta-D-glucanases, similar to Phanerochaete chrysosporium laminarinase 16A. Lam16A belongs to the 'nonspecific' 1,3(4)-beta-glucanase subfamily, although beta-1,6 branching and beta-1,4 bonds specifically define where Lam16A hydrolyzes its substrates, like curdlan (beta-1,3-glucan), lichenin (beta-1,3-1,4-mixed linkage glucan), and laminarin (beta-1,6-branched-1,3-glucan). |
| 185693 | GH16_laminarinase_like | 1.95e-09 | 156 | 289 | 52 | 166 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| 185691 | GH16_Strep_laminarinase_like | 8.35e-05 | 196 | 233 | 106 | 148 | Streptomyces laminarinase-like, member of glycosyl hydrolase family 16. Proteins similar to Streptomyces sioyaensis beta-1,3-glucanase (laminarinase) present in Actinomycetales as well as Peziomycotina. Laminarinases belong to glycosyl hydrolase family 16 and hydrolyze the glycosidic bond of the 1,3-beta-linked glucan, a major component of fungal and plant cell walls and the structural and storage polysaccharides (laminarin) of marine macro-algae. Members of the GH16 family have a conserved jelly roll fold with an active site channel. |
| 185694 | GH16_CCF | 0.010 | 198 | 233 | 130 | 179 | Coelomic cytolytic factor, member of glycosyl hydrolase family 16. Subgroup of glucanases of unknown function that are related to beta-GRP (beta-1,3-glucan recognition protein), but contain active site residues. Beta-GRPs are one group of pattern recognition receptors (PRRs), also referred to as biosensor proteins, that complexes with pathogen-associated beta-1,3-glucans and then transduces signals necessary for activation of an appropriate innate immune response. Beta-GRPs are present in insects and lack all catalytic residues. This subgroup contains related proteins that still contain the active site and are widely distributed in eukaryotes. Their structures adopt a jelly roll fold with a deep active site channel harboring the catalytic residues, like those of other glycosyl hydrolase family 16 members. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.76e-194 | 1 | 396 | 1 | 395 | |
| 9.90e-194 | 1 | 396 | 28 | 422 | |
| 1.10e-193 | 1 | 396 | 31 | 425 | |
| 1.10e-193 | 1 | 396 | 31 | 425 | |
| 8.78e-193 | 1 | 396 | 1 | 395 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.19e-84 | 104 | 394 | 3 | 298 | Chain A, PUTATIVE LAMINARINASE [Phanerodontia chrysosporium],2W39_A Chain A, Putative Laminarinase [Phanerodontia chrysosporium],2W52_A Chain A, PUTATIVE LAMINARINASE [Phanerodontia chrysosporium] |
|
| 4.06e-83 | 104 | 394 | 3 | 298 | Chain A, PUTATIVE LAMINARINASE [Phanerodontia chrysosporium],2WNE_A Chain A, PUTATIVE LAMINARINASE [Phanerodontia chrysosporium] |
|
| 8.81e-67 | 104 | 393 | 3 | 297 | Crystal structure and characterization an elongating GH family 16 beta-1,3-glucosyltransferase [Paecilomyces sp. 'thermophila'],5JVV_B Crystal structure and characterization an elongating GH family 16 beta-1,3-glucosyltransferase [Paecilomyces sp. 'thermophila'] |
|
| 6.73e-66 | 104 | 393 | 2 | 295 | The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_B The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_C The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_D The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'] |
|
| 6.93e-66 | 104 | 393 | 3 | 296 | The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_B The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_C The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_D The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDV_A The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_B The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_C The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_D The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.95e-87 | 95 | 394 | 120 | 418 | Probable glycosidase C21B10.07 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC21B10.07 PE=3 SV=1 |
|
| 6.75e-59 | 104 | 395 | 31 | 327 | Probable endo-1,3(4)-beta-glucanase AO090023000083 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=AO090023000083 PE=3 SV=1 |
|
| 9.39e-59 | 104 | 395 | 31 | 327 | Probable endo-1,3(4)-beta-glucanase AFLA_105200 OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=AFLA_105200 PE=3 SV=1 |
|
| 1.85e-58 | 104 | 394 | 20 | 315 | Endo-1,3(4)-beta-glucanase ARB_04519 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_04519 PE=3 SV=1 |
|
| 2.00e-58 | 104 | 394 | 32 | 327 | Probable endo-1,3(4)-beta-glucanase AFUB_029980 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=AFUB_029980 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
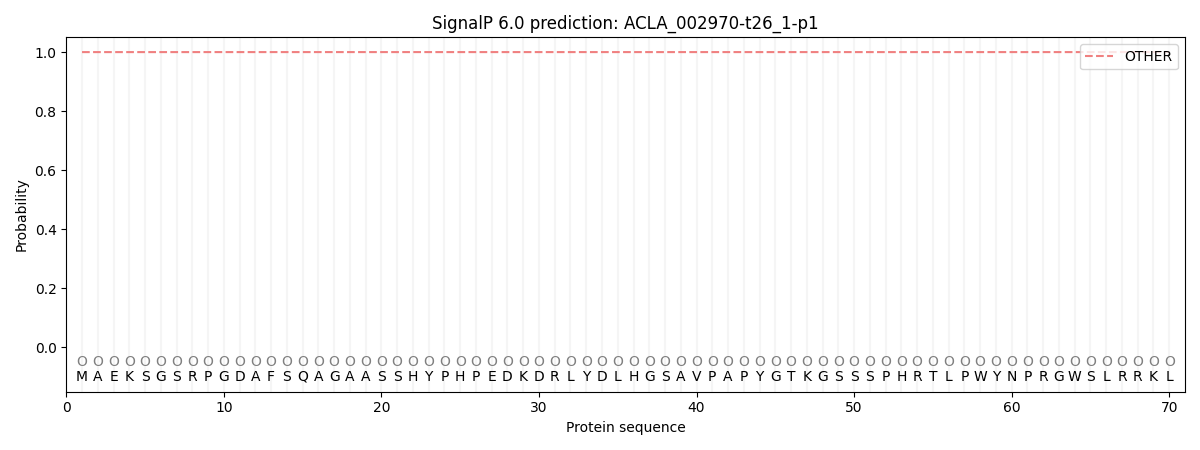
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000028 | 0.000008 |