Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS198 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN039 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF07728.19,PF08406.15,PF07726.16,PF00004.34,PF04851.20 | |||||||||||||
| Phrog | phrog_249 | |||||||||||||
| Host taxa | ;;;;;; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS166 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.78 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A0H3H502-F1-model_v4 | 65 | 366 | 1 | 281 | 0.9391 | 0.824 |
| AF-Q51481-F1-model_v4 | 93 | 366 | 1 | 260 | 0.9239 | 0.7833 |
| AF-Q2G2J8-F1-model_v4 | 100 | 365 | 1 | 263 | 0.9064 | 0.772 |
| AF-K0EW51-F1-model_v4 | 27 | 367 | 1 | 294 | 0.7878 | 0.7008 |
| AF-X8F327-F1-model_v4 | 64 | 367 | 5 | 289 | 0.7931 | 0.7002 |
| AF-P71922-F1-model_v4 | 24 | 367 | 1 | 291 | 0.7905 | 0.6998 |
| AF-O53705-F1-model_v4 | 66 | 367 | 1 | 297 | 0.7798 | 0.6974 |
| AF-K0FAA2-F1-model_v4 | 65 | 367 | 2 | 288 | 0.7837 | 0.6955 |
| AF-Q9I0D5-F1-model_v4 | 68 | 363 | 1 | 276 | 0.7762 | 0.6759 |
| AF-P33348-F1-model_v4 | 64 | 367 | 35 | 355 | 0.6858 | 0.6507 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP000089807778.pdb | 64 | 366 | 1 | 309 | 0.933 | 0.8546 |
| MGYP000300022970.pdb | 64 | 367 | 1 | 309 | 0.9267 | 0.8478 |
| MGYP000858003990.pdb | 67 | 366 | 1 | 306 | 0.9274 | 0.8453 |
| MGYP003663999631.pdb | 51 | 367 | 1 | 308 | 0.9216 | 0.8439 |
| MGYP001466739184.pdb | 57 | 366 | 3 | 319 | 0.9083 | 0.8433 |
| MGYP001626237157.pdb | 64 | 366 | 1 | 312 | 0.9137 | 0.8416 |
| MGYP001557083938.pdb | 59 | 367 | 1 | 312 | 0.907 | 0.8403 |
| MGYP003646054441.pdb | 58 | 367 | 1 | 315 | 0.9017 | 0.84 |
| MGYP001281949795.pdb | 58 | 367 | 1 | 321 | 0.8986 | 0.8376 |
| MGYP001467461195.pdb | 21 | 367 | 1 | 324 | 0.8947 | 0.8366 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS198 | IMGVR_UViG_3300050581_000016|3300050581|Ga0531113_000653_18870_19973 | 5 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
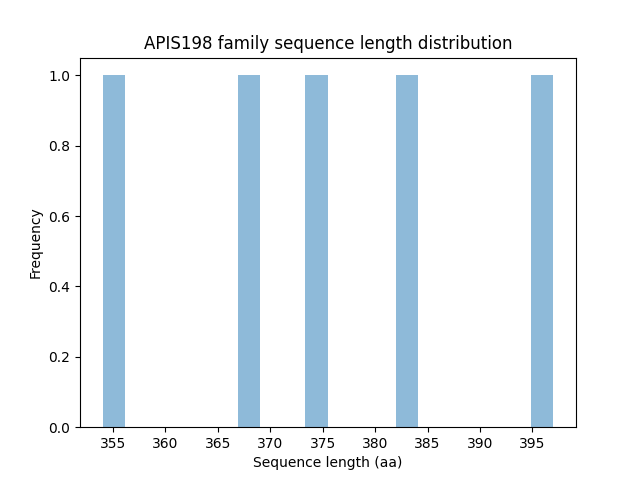
Family members, sources, and their hosts help
| Member | pident | evalue | bitscore | qcovs | phage | host | SourceDB | Pfam | Phrog |
|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_3300050581_000016|3300050581|Ga0531113_000653_18870_19973 | 100.000 | 4.029E-248 | 754 | 1.000% | IMGVR | PF07728 PF08406 PF07726 PF00004 PF04851 | phrog_249 | ||
| IMGVR_UViG_3300028388_005700|3300028388|Ga0307031_100107811 | 46.300 | 2.757E-91 | 301 | 0.973% | IMGVR | PF07728 PF08406 PF07726 PF00004 PF13191 | phrog_249 | ||
| IMGVR_UViG_3300028549_000664|3300028549|Ga0308317_14205878 | 49.400 | 2.635E-105 | 342 | 0.981% | IMGVR | PF07728 PF07726 PF08406 | phrog_249 | ||
| IMGVR_UViG_3300031111_000001|3300031111|Ga0272444_10000091132 | 46.600 | 1.204E-92 | 305 | 0.975% | IMGVR | PF07728 PF08406 PF07726 | phrog_249 | ||
| IMGVR_UViG_3300034685_000238|3300034685|Ga0310149_001011_22781_23974 | 47.600 | 2.289E-95 | 313 | 0.975% | IMGVR | PF07728 PF07726 PF00004 PF12775 | phrog_249 |
