Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS127 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN039 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF07728.19,PF07726.16,PF00004.34,PF08406.15,PF13173.11,PF13401.11 | |||||||||||||
| Phrog | phrog_249 | |||||||||||||
| Host taxa | ;;;;;; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS166 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.61 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A0H3H502-F1-model_v4 | 206 | 549 | 1 | 281 | 0.9461 | 0.7103 |
| AF-P33348-F1-model_v4 | 198 | 555 | 30 | 362 | 0.7274 | 0.5954 |
| AF-K0FCV8-F1-model_v4 | 198 | 556 | 25 | 372 | 0.7039 | 0.5814 |
| AF-K0EYL3-F1-model_v4 | 198 | 555 | 31 | 366 | 0.7028 | 0.577 |
| AF-A0A0G2K3W1-F1-model_v4 | 135 | 556 | 1 | 454 | 0.5751 | 0.5303 |
| AF-A0A132YZV7-F1-model_v4 | 233 | 551 | 1 | 389 | 0.6251 | 0.4175 |
| AF-A0A3P7PBI1-F1-model_v4 | 37 | 556 | 423 | 877 | 0.5213 | 0.4164 |
| AF-A0A0N4UFG6-F1-model_v4 | 36 | 556 | 231 | 739 | 0.474 | 0.3938 |
| AF-Q9H799-F1-model_v4 | 190 | 556 | 60 | 908 | 0.5214 | 0.3644 |
| AF-Q9HC84-F17-model_v4 | 37 | 556 | 593 | 1313 | 0.5193 | 0.3628 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP003686209621.pdb | 186 | 543 | 1 | 345 | 0.9447 | 0.763 |
| MGYP000605860649.pdb | 217 | 549 | 1 | 319 | 0.9516 | 0.7463 |
| MGYP001014096815.pdb | 228 | 552 | 1 | 323 | 0.9452 | 0.7445 |
| MGYP001189287878.pdb | 220 | 549 | 1 | 324 | 0.9337 | 0.7408 |
| MGYP003663999631.pdb | 215 | 553 | 1 | 311 | 0.9524 | 0.7403 |
| MGYP001269374496.pdb | 216 | 553 | 5 | 332 | 0.93 | 0.7401 |
| MGYP001588267429.pdb | 134 | 550 | 4 | 378 | 0.884 | 0.7379 |
| MGYP001352132134.pdb | 215 | 551 | 5 | 318 | 0.9383 | 0.7353 |
| MGYP000612083078.pdb | 214 | 555 | 3 | 323 | 0.9305 | 0.7329 |
| MGYP003706907671.pdb | 215 | 543 | 10 | 328 | 0.9244 | 0.7325 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS127 | IMGVR_UViG_3300006925_000049|3300006925|Ga0098050_100007221 | 7 | HMM model | Member alignment |
|---|
Host Taxa distribution
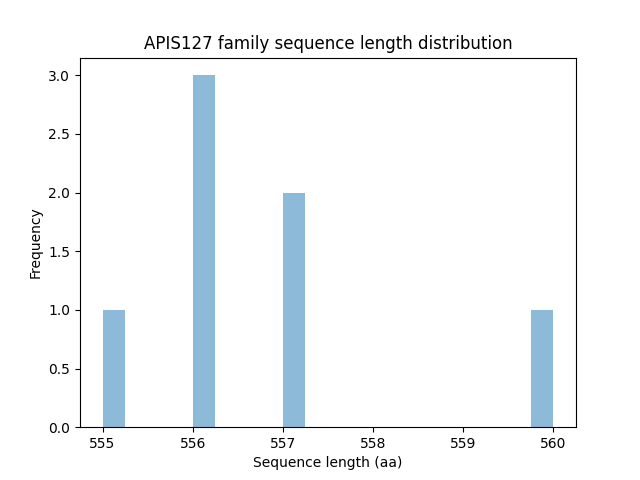
Length distribution