Seed proteins |
Seed genomic context | Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS115 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN049 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF00293.33,PF01467.31,PF00293.33,PF01467.31 | |||||||||||||
| Phrog | phrog_1185,phrog_1185 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Lysobacterales;f__Lysobacteraceae;g__Xanthomonas;s__unclassified Xanthomonas species | |||||||||||||
| Gene Location | Start: 122606; End: 123709; Strand: - | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending NAD+ reconstitution pathway (NARP) because it belongs to the same clan as family APIS170 which has seed protein. | |||||||||||||
Seed genomic context Download table help
| Genome accession | Start | End | Strand | Protein ID | Length | Molecular weight | Charge | Isoelectric point | Function | Phrog | PFAM |
|---|---|---|---|---|---|---|---|---|---|---|---|
| MN047793 | 120313 | 121452 | - | MN047793_00252 | 379 | 43401.17 | -2.5 | 5.6866 | hypothetical protein | phrog_4182 | |
| MN047793 | 121449 | 121613 | - | MN047793_00253 | 54 | 6179.06 | -1.0 | 5.6013 | hypothetical protein | ||
| MN047793 | 121610 | 122011 | - | MN047793_00254 | 133 | 14964.64 | -3.0 | 4.7920 | hypothetical protein | ||
| MN047793 | 122004 | 122282 | - | MN047793_00255 | 92 | 10655.00 | -6.5 | 4.3296 | hypothetical protein | ||
| MN047793 | 122350 | 122598 | - | MN047793_00256 | 82 | 9945.13 | -0.5 | 5.8075 | hypothetical protein | ||
| MN047793 | 122606 | 123709 | - | MN047793_00257 | 367 | 41914.66 | -3.0 | 5.9068 | Bifunctional NMN adenylyltransferase/Nudix hydrolase | phrog_1185 ; phrog_1185 | PF00293.33 ;PF01467.31 ;PF00293.33 ;PF01467.31 |
| MN047793 | 123799 | 124134 | + | MN047793_00258 | 111 | 12501.43 | 1.5 | 7.3201 | hypothetical protein | ||
| MN047793 | 124086 | 124301 | - | MN047793_00259 | 71 | 7584.84 | -1.0 | 4.8532 | hypothetical protein | PF04324.20 | |
| MN047793 | 124365 | 125498 | - | MN047793_00260 | 377 | 42253.62 | -19.0 | 4.5953 | hypothetical protein | ||
| MN047793 | 125509 | 125844 | - | MN047793_00261 | 111 | 12923.02 | 4.5 | 9.9083 | hypothetical protein | ||
| MN047793 | 125841 | 126230 | - | MN047793_00262 | 129 | 14525.76 | 1.0 | 7.9404 | hypothetical protein |
Seed genomic context- JBrowse
Predicted 3D structure by alphafold2 with pTM = 0.88 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-Q58361-F1-model_v4 | 9 | 366 | 48 | 338 | 0.5165 | 0.4947 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP000400551823.pdb | 6 | 367 | 1 | 340 | 0.9709 | 0.9304 |
| MGYP001592028481.pdb | 7 | 367 | 1 | 341 | 0.967 | 0.9272 |
| MGYP001111241452.pdb | 9 | 367 | 1 | 344 | 0.9618 | 0.927 |
| MGYP003574359643.pdb | 5 | 367 | 7 | 351 | 0.9518 | 0.9264 |
| MGYP000138452357.pdb | 6 | 366 | 1 | 344 | 0.9598 | 0.9242 |
| MGYP001390947301.pdb | 6 | 366 | 1 | 350 | 0.9475 | 0.9202 |
| MGYP001822565791.pdb | 1 | 367 | 1 | 350 | 0.948 | 0.9198 |
| MGYP002652398978.pdb | 2 | 367 | 4 | 348 | 0.9487 | 0.9183 |
| MGYP003652350255.pdb | 8 | 366 | 1 | 342 | 0.9554 | 0.9165 |
| MGYP001371600575.pdb | 6 | 366 | 1 | 351 | 0.9414 | 0.9155 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS115 | MN047793_00257 | 6 | HMM model | Member alignment |
|---|
Host Taxa distribution
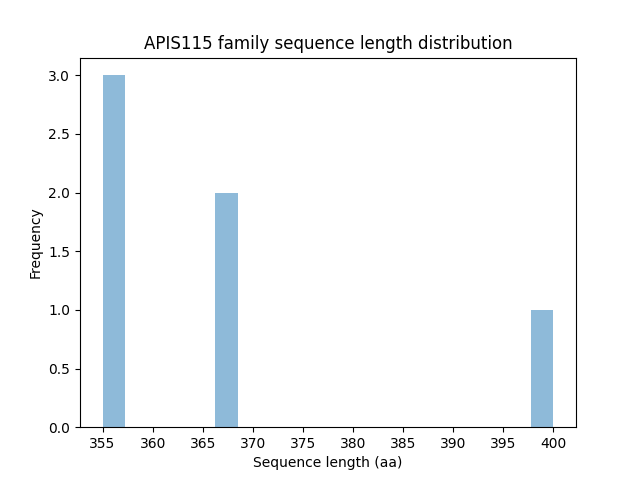
Length distribution