Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS106 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN044 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | ||||||||||||||
| Phrog | phrog_6408 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Enterobacterales;f__Erwiniaceae;g__Erwinia;s__unclassified Erwinia species | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending TIR-STING because it belongs to the same clan as family APIS150 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.51 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
No homologs found in esmfold database
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS106 | OP480062_00009 | 4 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
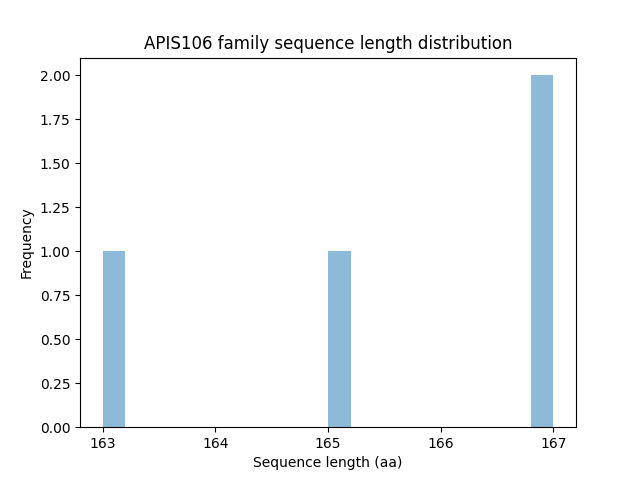
Family members, sources, and their hosts help
| Member | pident | evalue | bitscore | qcovs | phage | host | SourceDB | Pfam | Phrog |
|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_3300048919_000048|3300048919|Ga0496116_0000137_143455_143952 | 51.700 | 4.561E-47 | 163 | 0.994% | IMGVR | phrog_6408 | |||
| OP480062_00009 | 100.000 | 7.788E-113 | 353 | 1.000% | Erwinia phage pEa_SNUABM_55 | Erwinia | INPHARED | phrog_6408 | |
| OP480063_00076 | 99.000 | 2.007E-112 | 351 | 1.000% | Erwinia phage pEa_SNUABM_56 | Erwinia | INPHARED | phrog_6408 | |
| MT552976_00098 | 49.000 | 3.662E-32 | 120 | 0.778% | Pectobacterium phage POP17 | Pectobacterium | INPHARED | phrog_6408 |
