Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS086 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN011 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | ||||||||||||||
| Phrog | phrog_8921,phrog_2375,phrog_3272,phrog_12748 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Enterobacterales;f__Enterobacteriaceae;g__Escherichia; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending toxin-antitoxin (TA) because it belongs to the same clan as family APIS012 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.58 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
No homologs found in esmfold database
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS086 | ON229909_00228 | 14 | HMM model | Member alignment |
|---|
Host Taxa distribution
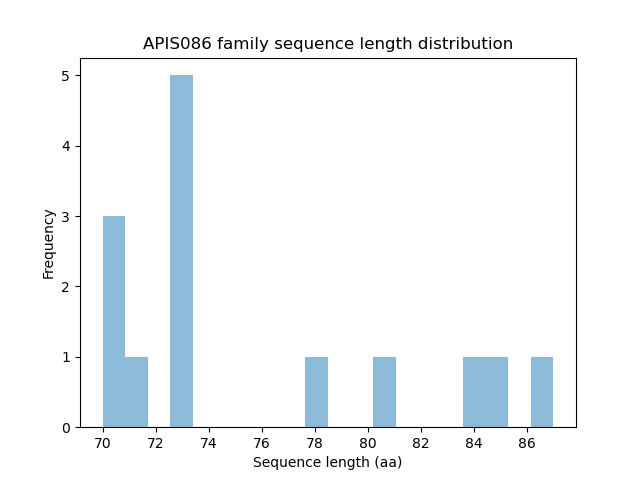
Length distribution
Homology to other families download full data without filtering help
Clan ID: CLAN011
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS012 | 1.81E-49 | 1 | 73 | 14 | 86 | toxin-antitoxin (TA) | NP_049652.1 |
Family members, sources, and their hosts help
| Member | pident | evalue | bitscore | qcovs | phage | host | SourceDB | Pfam | Phrog |
|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_2974743505_000001|2974743505|2974743655 | 100.000 | 3.09e-46 | 150 | 84% | IMGVR | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |||
| MGV-GENOME-0377451_82 | 100.000 | 1.68e-46 | 150 | 90% | MGV | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |||
| MGV-GENOME-0377517_58 | 57.143 | 2.53e-20 | 84.7 | 99% | MGV | phrog_8921 phrog_2375 phrog_12748 | |||
| MK327943_00035 | 55.714 | 1.59e-19 | 82.4 | 100% | Escherichia phage vB_EcoM_G53 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| MK373782_00037 | 57.143 | 6.97e-20 | 83.6 | 100% | Escherichia phage vB_EcoM_KAW3E185 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_12748 | |
| MK886800_00269 | 98.630 | 9.24e-46 | 149 | 100% | Escherichia phage EcNP1 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| MW883059_00026 | 98.630 | 7.20e-45 | 146 | 100% | Escherichia phage vB_EcoM_SYGD1 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| MZ189262_00034 | 83.117 | 2.41e-37 | 128 | 92% | Escherichia phage W143 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| MZ502380_00035 | 97.260 | 1.34e-45 | 148 | 100% | Escherichia phage vB_EcoM_Nami | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| ON229909_00228 | 100.000 | 2.73e-46 | 150 | 100% | Escherichia phage vB_EcoM_CE1 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| ON506924_00037 | 98.630 | 1.61e-45 | 148 | 100% | Escherichia phage vB_EcoM_SCS4 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| ON720975_00159 | 98.630 | 4.33e-46 | 149 | 100% | Salmonella phage GRNsp7 | Salmonella | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| OP172786_00049 | 95.890 | 3.62e-44 | 145 | 100% | Escherichia phage ECO07P3 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_3272 phrog_12748 | |
| OQ703618_00189 | 57.143 | 2.54e-20 | 84.7 | 96% | Escherichia phage GADS24 | Escherichia | INPHARED | phrog_8921 phrog_2375 phrog_12748 |
| APIS | Contig | Range |
|---|---|---|
| APIS086 | - |
