Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS080 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN010 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF17574.5 | |||||||||||||
| Phrog | phrog_2254 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Enterobacterales;f__Enterobacteriaceae;g__Klebsiella; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending toxin-antitoxin (TA) because it belongs to the same clan as family APIS011 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.73 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP003595835577.pdb | 8 | 96 | 3 | 89 | 0.7983 | 0.7126 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS080 | ON602753_00020 | 39 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
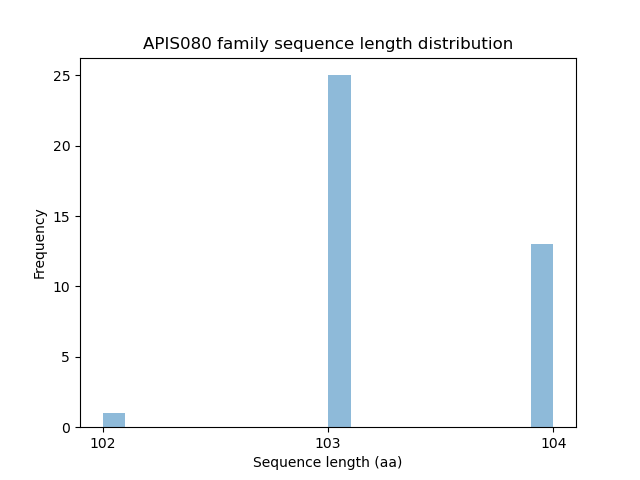
Homology to other families download full data without filtering help
Clan ID: CLAN010
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS011 | 5.48E-32 | 9 | 86 | 26 | 103 | toxin-antitoxin (TA) | NP_041980.1 |
| APIS084 | 8.32E-26 | 6 | 77 | 7 | 78 | - | MW001769_00023 |
| APIS078 | 2.51E-23 | 9 | 74 | 5 | 70 | - | KX078568_00026 |
Family members, sources, and their hosts help
| Member | pident | evalue | bitscore | qcovs | phage | host | SourceDB | Pfam | Phrog |
|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_3300045988_000298|3300045988|Ga0495776_113816_26626_26937 | 92.857 | 2.15e-60 | 188 | 95% | IMGVR | PF17574 | phrog_2254 | ||
| IMGVR_UViG_3300045988_001943|3300045988|Ga0495776_046194_7165_7476 | 88.119 | 1.81e-58 | 183 | 98% | IMGVR | PF17574 | phrog_2254 | ||
| MGV-GENOME-0149923_11 | 92.157 | 3.76e-60 | 187 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0189393_13 | 88.119 | 1.89e-58 | 183 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0265429_14 | 93.069 | 4.12e-62 | 192 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0266883_34 | 90.099 | 1.53e-59 | 186 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0269628_21 | 94.175 | 7.61e-63 | 194 | 98% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0271769_13 | 91.000 | 1.32e-60 | 188 | 96% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0271873_37 | 92.157 | 6.42e-61 | 189 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0271896_27 | 88.119 | 3.00e-58 | 182 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0272093_12 | 92.857 | 2.24e-60 | 188 | 94% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0272520_27 | 89.320 | 1.99e-59 | 185 | 98% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0274788_15 | 91.176 | 2.37e-60 | 187 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0275239_15 | 92.079 | 6.50e-60 | 187 | 97% | MGV | PF17574 | phrog_2254 | ||
| MGV-GENOME-0276411_33 | 89.109 | 6.74e-59 | 184 | 97% | MGV | PF17574 | phrog_2254 | ||
| MH059637_00020 | 70.370 | 9.81e-41 | 121 | 79% | Pectobacterium phage Jarilo | Pectobacterium | INPHARED | PF17574 | phrog_2254 |
| MK795385_00033 | 94.175 | 7.30e-63 | 194 | 99% | Klebsiella phage IME304 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| MT813141_00017 | 92.929 | 1.67e-60 | 188 | 96% | Klebsiella phage NL_ZS_2 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| MT966872_00022 | 90.000 | 5.23e-60 | 187 | 97% | Klebsiella phage P510 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| MW042791_00025 | 90.722 | 1.64e-58 | 183 | 94% | Klebsiella phage 066023 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| MW042800_00047 | 91.000 | 1.27e-60 | 188 | 97% | Klebsiella phage 066037 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| MW042812_00012 | 90.196 | 5.35e-60 | 187 | 98% | Klebsiella phage 066134 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OM194189_00019 | 93.069 | 3.96e-62 | 192 | 98% | Klebsiella phage Shaphc-9226R | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| ON602753_00020 | 100.000 | 1.08e-67 | 206 | 100% | Klebsiella phage VLCpiA3d | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| ON602755_00024 | 91.000 | 8.58e-60 | 186 | 97% | Klebsiella phage VLCpiA3c | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| ON880501_00029 | 92.079 | 1.19e-61 | 191 | 98% | Klebsiella phage vB_KpnP1 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OP256047_00028 | 93.137 | 2.09e-54 | 172 | 99% | Klebsiella phage BUCT-3589 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OQ579029_00019 | 92.000 | 9.70e-54 | 171 | 97% | Klebsiella phage Saitama | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OQ579030_00020 | 89.109 | 9.52e-52 | 166 | 98% | Klebsiella phage Emom | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OX335390_00021 | 87.129 | 2.67e-50 | 162 | 98% | Klebsiella phage cp7 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| OX335398_00022 | 88.119 | 6.15e-51 | 164 | 98% | Klebsiella phage cp8 | Klebsiella | INPHARED | PF17574 | phrog_2254 |
| uvig_225801_27 | 89.320 | 1.91e-59 | 185 | 99% | GPD | PF17574 | phrog_2254 | ||
| uvig_234063_32 | 88.119 | 2.88e-58 | 182 | 98% | GPD | PF17574 | phrog_2254 | ||
| uvig_241747_16 | 93.137 | 1.35e-61 | 191 | 98% | GPD | PF17574 | phrog_2254 | ||
| uvig_282596_29 | 98.058 | 1.69e-64 | 198 | 99% | GPD | PF17574 | phrog_2254 | ||
| uvig_320142_31 | 91.176 | 2.27e-60 | 187 | 98% | GPD | PF17574 | phrog_2254 | ||
| uvig_325023_26 | 92.079 | 6.24e-60 | 187 | 98% | GPD | PF17574 | phrog_2254 | ||
| uvig_328834_17 | 90.099 | 1.47e-59 | 186 | 98% | GPD | PF17574 | phrog_2254 | ||
| uvig_472708_11 | 92.157 | 3.60e-60 | 187 | 98% | GPD | PF17574 | phrog_2254 |
| APIS | Contig | Range |
|---|---|---|
| APIS080 | - |
