Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS008 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | Gabija | |||||||||||||
| CLAN ID | CLAN007 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | Yirmiya et al., 2023 | |||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | ||||||||||||||
| Phrog | phrog_18122,phrog_27100,phrog_4400,phrog_25514,phrog_37109,phrog_16176 | |||||||||||||
| Host taxa | d__Bacteria;p__Firmicutes;c__Bacilli;o__Bacillales;f__Bacillaceae;g__Bacillus;s__Bacillus subtilis | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | Alphafold2 predicted that Gad2 is an enzyme with a nucleotidyltransferase protein domain, suggesting that it inhibits Gabija via a mechanism of action different than Gad1. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.71 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP000373333736.pdb | 1 | 221 | 1 | 204 | 0.9299 | 0.7 |
| MGYP003394315973.pdb | 1 | 199 | 2 | 202 | 0.9177 | 0.6875 |
| MGYP000999235125.pdb | 3 | 220 | 1 | 204 | 0.912 | 0.6868 |
| MGYP001439174798.pdb | 1 | 229 | 1 | 230 | 0.8785 | 0.6862 |
| MGYP003537377462.pdb | 2 | 400 | 1 | 266 | 0.8166 | 0.6752 |
| MGYP000376116683.pdb | 1 | 264 | 2 | 248 | 0.8319 | 0.6687 |
| MGYP000981939358.pdb | 1 | 235 | 2 | 199 | 0.8876 | 0.6603 |
| MGYP001240424264.pdb | 1 | 228 | 2 | 229 | 0.8384 | 0.6572 |
| MGYP001238726922.pdb | 1 | 219 | 1 | 226 | 0.871 | 0.6473 |
| MGYP001760742232.pdb | 12 | 223 | 1 | 235 | 0.8161 | 0.6428 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS008 | Gad2 | 49 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
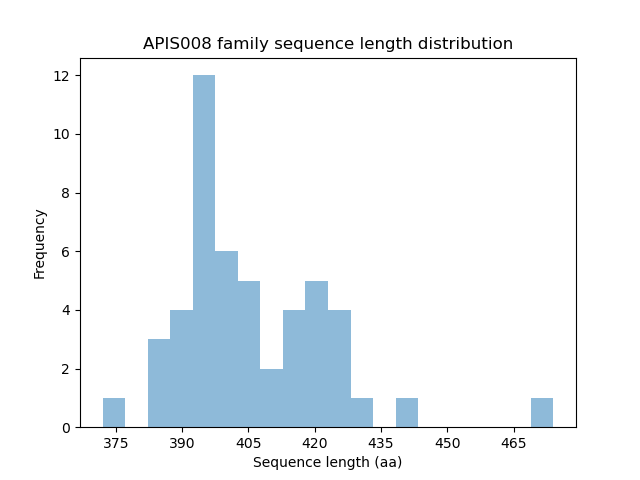
Homology to other families download full data without filtering help
Clan ID: CLAN007
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS056 | 2.00E-77 | 75 | 468 | 2 | 364 | - | IMGVR_UViG_3300020049_002891|3300020049|Ga0206652_100507913 |
| APIS059 | 7.46E-67 | 58 | 457 | 2 | 399 | - | KF302034_00153 |
| APIS089 | 1.00E-51 | 77 | 458 | 2 | 389 | - | uvig_330502_68 |
