You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004770_01557
You are here: Home > Sequence: MGYG000004770_01557
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Streptococcus sobrinus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus sobrinus | |||||||||||
| CAZyme ID | MGYG000004770_01557 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14711; End: 15352 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3942 | COG3942 | 2.27e-24 | 85 | 183 | 47 | 149 | Surface antigen [Cell wall/membrane/envelope biogenesis]. |
| pfam05257 | CHAP | 3.98e-18 | 103 | 181 | 2 | 83 | CHAP domain. This domain corresponds to an amidase function. Many of these proteins are involved in cell wall metabolism of bacteria. This domain is found at the N-terminus of Escherichia coli gss, where it functions as a glutathionylspermidine amidase EC:3.5.1.78. This domain is found to be the catalytic domain of PlyCA. CHAP is the amidase domain of bifunctional Escherichia coli glutathionylspermidine synthetase/amidase, and it catalyzes the hydrolysis of Gsp (glutathionylspermidine) into glutathione and spermidine. |
| PRK08581 | PRK08581 | 6.64e-16 | 102 | 212 | 504 | 619 | amidase domain-containing protein. |
| cd00118 | LysM | 2.46e-06 | 29 | 72 | 2 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| pfam01476 | LysM | 4.80e-05 | 30 | 70 | 1 | 40 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWN62848.1 | 7.17e-146 | 1 | 213 | 1 | 213 |
| SQG18783.1 | 7.17e-146 | 1 | 213 | 1 | 213 |
| AWN18203.1 | 7.17e-146 | 1 | 213 | 1 | 213 |
| AWN20123.1 | 7.17e-146 | 1 | 213 | 1 | 213 |
| SQG12836.1 | 7.17e-146 | 1 | 213 | 1 | 213 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4CGK_A | 1.62e-33 | 103 | 210 | 284 | 390 | Crystalstructure of the essential protein PcsB from Streptococcus pneumoniae [Streptococcus pneumoniae D39],4CGK_B Crystal structure of the essential protein PcsB from Streptococcus pneumoniae [Streptococcus pneumoniae D39] |
| 2K3A_A | 2.51e-19 | 88 | 186 | 29 | 133 | ChainA, CHAP domain protein [Staphylococcus saprophyticus subsp. saprophyticus ATCC 15305 = NCTC 7292] |
| 2LRJ_A | 2.45e-13 | 97 | 186 | 2 | 92 | ChainA, Staphyloxanthin biosynthesis protein, putative [Staphylococcus aureus subsp. aureus COL] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8DVU8 | 1.26e-72 | 5 | 212 | 6 | 211 | Putative hydrolase SMU_367 OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=SMU_367 PE=3 SV=1 |
| Q2G0D4 | 1.45e-23 | 23 | 192 | 84 | 251 | Probable autolysin SsaALP OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=SAOUHSC_00671 PE=1 SV=1 |
| Q49UX4 | 1.69e-16 | 29 | 210 | 151 | 325 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus saprophyticus subsp. saprophyticus (strain ATCC 15305 / DSM 20229 / NCIMB 8711 / NCTC 7292 / S-41) OX=342451 GN=sle1 PE=3 SV=1 |
| Q5HCY4 | 2.62e-15 | 50 | 182 | 98 | 231 | Staphylococcal secretory antigen ssaA1 OS=Staphylococcus aureus (strain COL) OX=93062 GN=ssaA1 PE=3 SV=1 |
| Q53587 | 2.62e-15 | 50 | 182 | 98 | 231 | Staphylococcal secretory antigen SsaA OS=Staphylococcus aureus (strain Newman) OX=426430 GN=ssaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
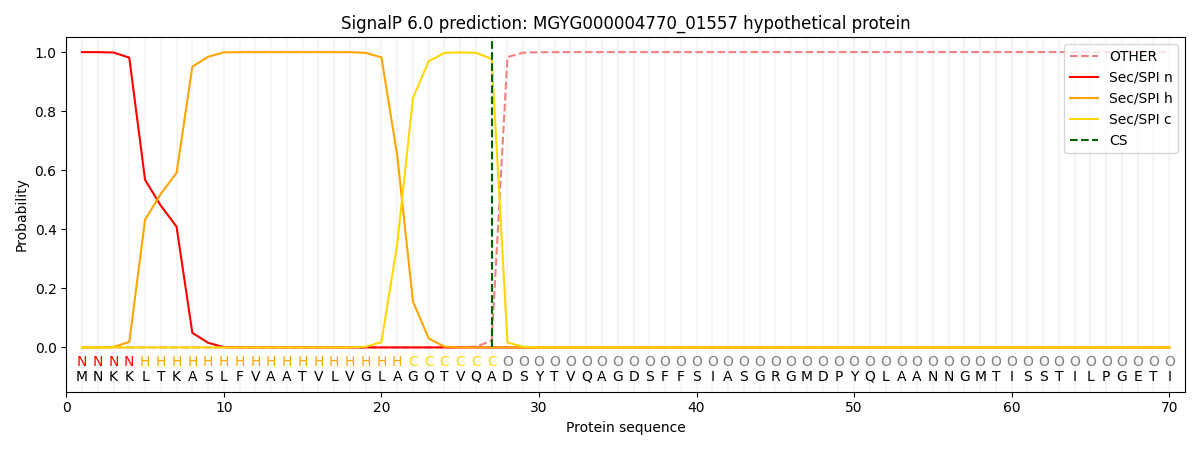
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000245 | 0.998937 | 0.000240 | 0.000187 | 0.000186 | 0.000158 |
