You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004707_01379
You are here: Home > Sequence: MGYG000004707_01379
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
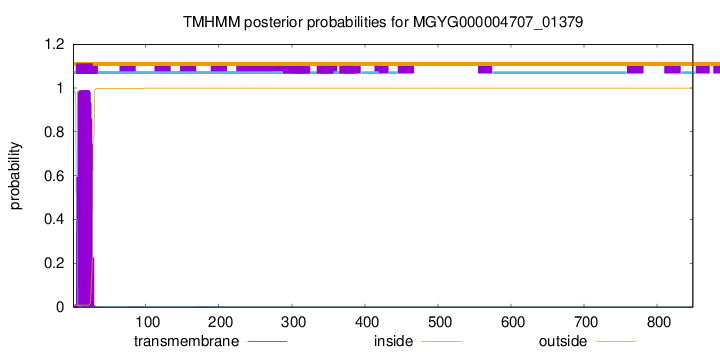
TMHMM annotations
Basic Information help
| Species | Monoglobus sp900554105 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; Monoglobus; Monoglobus sp900554105 | |||||||||||
| CAZyme ID | MGYG000004707_01379 | |||||||||||
| CAZy Family | CE12 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 323; End: 2872 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE12 | 574 | 818 | 9.5e-53 | 0.9952380952380953 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01821 | Rhamnogalacturan_acetylesterase_like | 6.27e-49 | 574 | 818 | 2 | 198 | Rhamnogalacturan_acetylesterase_like subgroup of SGNH-hydrolases. Rhamnogalacturan acetylesterase removes acetyl esters from rhamnogalacturonan substrates, and renders them susceptible to degradation by rhamnogalacturonases. Rhamnogalacturonans are highly branched regions in pectic polysaccharides, consisting of repeating -(1,2)-L-Rha-(1,4)-D-GalUA disaccharide units, with many rhamnose residues substituted by neutral oligosaccharides such as arabinans, galactans and arabinogalactans. Extracellular enzymes participating in the degradation of plant cell wall polymers, such as Rhamnogalacturonan acetylesterase, would typically be found in saprophytic and plant pathogenic fungi and bacteria. |
| COG2755 | TesA | 4.98e-11 | 567 | 733 | 3 | 172 | Lysophospholipase L1 or related esterase [Amino acid transport and metabolism]. |
| cd00229 | SGNH_hydrolase | 5.46e-06 | 575 | 753 | 1 | 181 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| pfam13472 | Lipase_GDSL_2 | 1.19e-05 | 579 | 733 | 3 | 148 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| cd01825 | SGNH_hydrolase_peri1 | 3.85e-04 | 605 | 738 | 12 | 149 | SGNH_peri1; putative periplasmic member of the SGNH-family of hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUO20372.1 | 0.0 | 1 | 845 | 5 | 865 |
| AUO20219.1 | 3.19e-52 | 575 | 819 | 125 | 351 |
| ATL46500.1 | 2.53e-33 | 572 | 825 | 24 | 241 |
| QII82135.1 | 2.93e-33 | 571 | 821 | 5 | 220 |
| APZ49497.1 | 3.99e-33 | 571 | 821 | 5 | 220 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5HO0_A | 9.74e-11 | 359 | 454 | 548 | 645 | Crystalstructure of AbnA (closed conformation), a GH43 extracellular arabinanase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],5HO2_A Crystal structure of AbnA (open conformation), a GH43 extracellular arabinanase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],5HOF_A Crystal structure of AbnA, a GH43 extracellular arabinanase from Geobacillus stearothermophilus, in complex with arabinopentaose [Geobacillus stearothermophilus],5HP6_A Structure of AbnA, a GH43 extracellular arabinanase from Geobacillus stearothermophilus (a new conformational state) [Geobacillus stearothermophilus] |
| 2O14_A | 1.28e-10 | 572 | 752 | 162 | 335 | X-RayCrystal Structure of Protein YXIM_BACsu from Bacillus subtilis. Northeast Structural Genomics Consortium Target SR595 [Bacillus subtilis] |
| 5HO9_A | 1.43e-10 | 359 | 448 | 548 | 639 | Structureof truncated AbnA (domains 1-3), a GH43 arabinanase from Geobacilllus stearothermophilus, in complex with arabinooctaose [Geobacillus stearothermophilus],5HO9_B Structure of truncated AbnA (domains 1-3), a GH43 arabinanase from Geobacilllus stearothermophilus, in complex with arabinooctaose [Geobacillus stearothermophilus] |
| 1DEO_A | 1.79e-07 | 574 | 657 | 2 | 78 | RHAMNOGALACTURONANACETYLESTERASE FROM ASPERGILLUS ACULEATUS AT 1.55 A RESOLUTION WITH SO4 IN THE ACTIVE SITE [Aspergillus aculeatus],1DEX_A RHAMNOGALACTURONAN ACETYLESTERASE FROM ASPERGILLUS ACULEATUS AT 1.9 A RESOLUTION [Aspergillus aculeatus],1K7C_A Rhamnogalacturonan acetylesterase with seven N-linked carbohydrate residues distributed at two N-glycosylation sites refined at 1.12 A resolution [Aspergillus aculeatus],1PP4_A The crystal structure of rhamnogalacturonan acetylesterase in space group P3121 [Aspergillus aculeatus],1PP4_B The crystal structure of rhamnogalacturonan acetylesterase in space group P3121 [Aspergillus aculeatus] |
| 3C1U_A | 1.79e-07 | 574 | 657 | 2 | 78 | ChainA, Rhamnogalacturonan acetylesterase [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O31523 | 3.75e-25 | 575 | 825 | 8 | 219 | Rhamnogalacturonan acetylesterase RhgT OS=Bacillus subtilis (strain 168) OX=224308 GN=rhgT PE=1 SV=1 |
| O31528 | 1.19e-20 | 575 | 752 | 5 | 174 | Probable rhamnogalacturonan acetylesterase YesY OS=Bacillus subtilis (strain 168) OX=224308 GN=yesY PE=1 SV=1 |
| P42304 | 9.95e-11 | 572 | 752 | 177 | 350 | Uncharacterized esterase YxiM OS=Bacillus subtilis (strain 168) OX=224308 GN=yxiM PE=1 SV=2 |
| Q5BAA2 | 1.53e-10 | 574 | 746 | 19 | 195 | Rhamnogalacturonan acetylesterase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=AN2528 PE=1 SV=1 |
| Q00017 | 1.18e-06 | 574 | 657 | 19 | 95 | Rhamnogalacturonan acetylesterase OS=Aspergillus aculeatus OX=5053 GN=rha1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
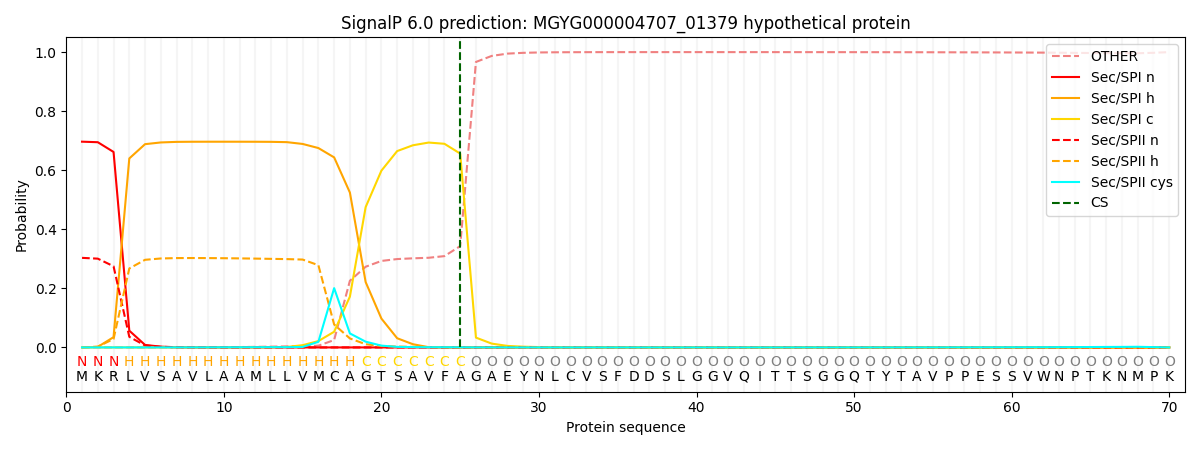
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001615 | 0.682624 | 0.314493 | 0.000574 | 0.000374 | 0.000306 |