You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004687_01628
You are here: Home > Sequence: MGYG000004687_01628
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Haemophilus_A parahaemolyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Pasteurellaceae; Haemophilus_A; Haemophilus_A parahaemolyticus | |||||||||||
| CAZyme ID | MGYG000004687_01628 | |||||||||||
| CAZy Family | GT32 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5561; End: 6343 Strand: + | |||||||||||
Full Sequence Download help
| MSIKYEKLIV LCNGLSRLLG NIVKILGYPF HALFPKKRFT IPEFSPAKLS SNRKDKINRT | 60 |
| IWQTNYSNKV TLPMYCNYLF NRLMSLSYDY RYVSTEEREE YLKTHAEPRV FEAYSQLTDG | 120 |
| AAQADFWRVF TLWREGGIYI DIDGHLVFPL SQIIRENDNE VLIKRRDKYT NFFLACEAGH | 180 |
| PIMKATLDRI VDNIENRRID GGVFVMTGPD TLNAAIGNQI VNTRRDKITC TQGTFTNEHF | 240 |
| QYIDKPRSKW NYKKNEELLK | 260 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3774 | OCH1 | 2.70e-63 | 3 | 260 | 10 | 289 | Mannosyltransferase OCH1 or related enzyme [Cell wall/membrane/envelope biogenesis]. |
| pfam04488 | Gly_transf_sug | 7.21e-18 | 73 | 158 | 1 | 91 | Glycosyltransferase sugar-binding region containing DXD motif. The DXD motif is a short conserved motif found in many families of glycosyltransferases, which add a range of different sugars to other sugars, phosphates and proteins. DXD-containing glycosyltransferases all use nucleoside diphosphate sugars as donors and require divalent cations, usually manganese. The DXD motif is expected to play a carbohydrate binding role in sugar-nucleoside diphosphate and manganese dependent glycosyltransferases. |
| cd16977 | VHS_GGA | 0.005 | 18 | 73 | 60 | 118 | VHS (Vps27/Hrs/STAM) domain of GGA (Golgi-localized, Gamma-ear-containing, Arf-binding) subfamily. GGA (Golgi-localized, Gamma-ear-containing, Arf-binding) comprises a subfamily of ubiquitously expressed, monomeric, motif-binding cargo/clathrin adaptor proteins involved in membrane trafficking between the Trans-Golgi Network (TGN) and endosomes. The VHS domain has a superhelical structure similar to the structure of the ARM (Armadillo) repeats and is present at the N-termini of proteins. GGA proteins have a multidomain structure consisting of an N-terminal VHS domain linked by a short proline-rich linker to a GAT (GGA and TOM) domain, which is followed by a long flexible linker to the C-terminal appendage, GAE (Gamma-Adaptin Ear) domain. The VHS domain of GGA proteins binds to the acidic-cluster dileucine (DxxLL) motif found on the cytoplasmic tails of cargo proteins trafficked between the Trans-Golgi Network and the endosomal system. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRP12239.1 | 3.43e-190 | 1 | 260 | 1 | 260 |
| QEN11046.1 | 3.43e-190 | 1 | 260 | 1 | 260 |
| AWI50249.1 | 2.94e-166 | 1 | 260 | 1 | 260 |
| SNV36599.1 | 8.85e-157 | 3 | 260 | 5 | 262 |
| AFU19707.1 | 8.85e-157 | 3 | 260 | 5 | 262 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P46393 | 7.27e-104 | 3 | 181 | 5 | 183 | Uncharacterized glycosyltransferase in aroQ 3'region OS=Actinobacillus pleuropneumoniae OX=715 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
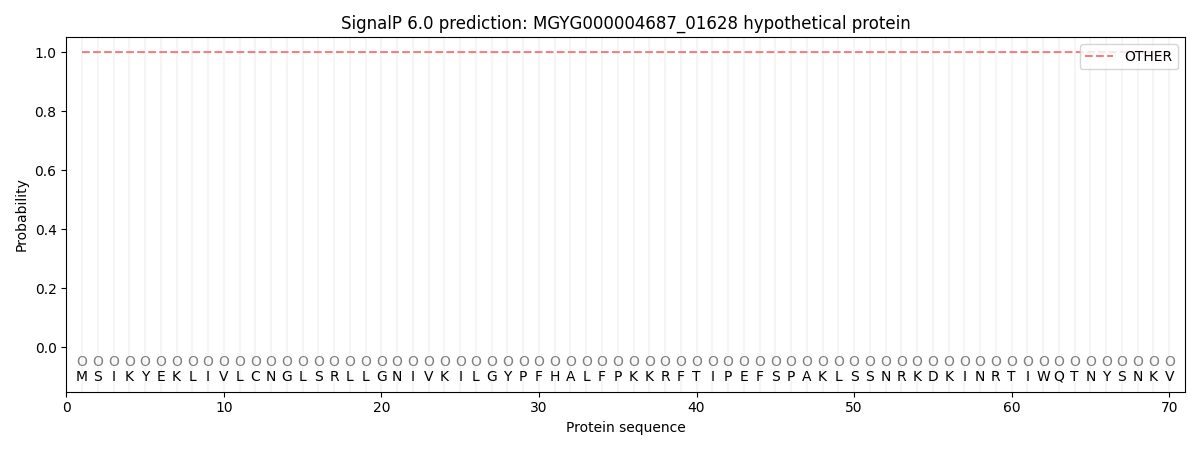
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000036 | 0.000009 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
