You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004345_00554
You are here: Home > Sequence: MGYG000004345_00554
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
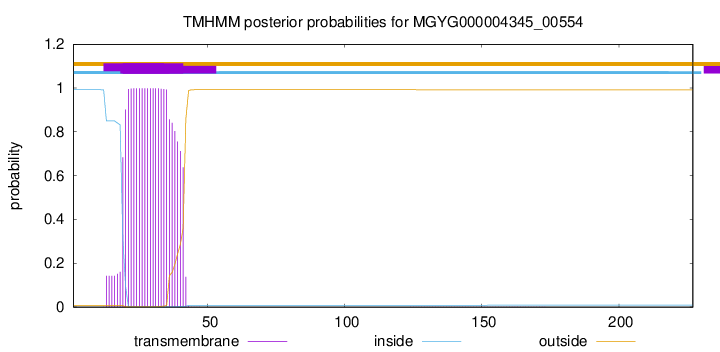
TMHMM annotations
Basic Information help
| Species | Streptococcus thermophilus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus thermophilus | |||||||||||
| CAZyme ID | MGYG000004345_00554 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 37719; End: 38402 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13925 | RPF | 1.53e-09 | 168 | 225 | 1 | 68 | core lysozyme-like domain of resuscitation-promoting factor proteins. Resuscitation-promoting factor (RPF) proteins, found in various (G+C)-rich Gram-positive bacteria, act to reactivate cultures from stationary phase. This protein shares elements of the structural core of lysozyme and related proteins. Furthermore, it shares a conserved active site glutamate which is required for activity, and has a polysaccharide binding cleft that corresponds to the peptidoglycan binding cleft of lysozyme. Muralytic activity of Rpf in Micrococcus luteus correlates with resuscitation, supporting a mechanism dependent on cleavage of peptidoglycan by RPF. |
| cd00254 | LT-like | 0.002 | 172 | 202 | 6 | 36 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd00442 | Lyz-like | 0.007 | 169 | 215 | 1 | 58 | lysozyme-like domains. This family contains several members, including soluble lytic transglycosylases (SLT), goose egg-white lysozymes (GEWL), hen egg-white lysozymes (HEWL), chitinases, bacteriophage lambda lysozymes, endolysins, autolysins, chitosanases, and pesticin. Typical members are involved in the hydrolysis of beta-1,4- linked polysaccharides. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATH75448.1 | 2.18e-131 | 1 | 227 | 1 | 227 |
| CAD0166394.1 | 2.18e-131 | 1 | 227 | 1 | 227 |
| QHD71358.1 | 2.18e-131 | 1 | 227 | 1 | 227 |
| CAD0166110.1 | 2.18e-131 | 1 | 227 | 1 | 227 |
| QKM73636.1 | 2.18e-131 | 1 | 227 | 1 | 227 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8CMZ9 | 1.30e-31 | 139 | 225 | 147 | 233 | Probable transglycosylase IsaA OS=Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200) OX=176280 GN=isaA PE=3 SV=1 |
| Q5HL49 | 1.30e-31 | 139 | 225 | 147 | 233 | Probable transglycosylase IsaA OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=isaA PE=3 SV=1 |
| Q6GDN1 | 6.60e-27 | 139 | 227 | 145 | 233 | Probable transglycosylase IsaA OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=isaA PE=3 SV=1 |
| Q2YWD9 | 2.56e-26 | 139 | 227 | 145 | 233 | Probable transglycosylase IsaA OS=Staphylococcus aureus (strain bovine RF122 / ET3-1) OX=273036 GN=isaA PE=3 SV=1 |
| Q6G6A5 | 9.89e-26 | 139 | 227 | 145 | 233 | Probable transglycosylase IsaA OS=Staphylococcus aureus (strain MSSA476) OX=282459 GN=isaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.394144 | 0.600988 | 0.002486 | 0.001101 | 0.000482 | 0.000792 |