You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004164_01523
You are here: Home > Sequence: MGYG000004164_01523
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1494 sp900552305 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; UMGS1494; UMGS1494 sp900552305 | |||||||||||
| CAZyme ID | MGYG000004164_01523 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | Anti-sigma-I factor RsgI6 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6262; End: 7590 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 53 | 382 | 3.6e-47 | 0.9735973597359736 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 1.68e-33 | 98 | 382 | 3 | 263 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 1.06e-25 | 98 | 384 | 69 | 339 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam00331 | Glyco_hydro_10 | 3.44e-23 | 78 | 384 | 24 | 310 | Glycosyl hydrolase family 10. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QMW04637.1 | 9.66e-107 | 19 | 432 | 32 | 436 |
| QMW04636.1 | 3.54e-98 | 19 | 436 | 35 | 451 |
| QHV94149.1 | 7.58e-97 | 19 | 442 | 32 | 445 |
| QIP12711.1 | 3.01e-96 | 19 | 442 | 32 | 445 |
| AXE20297.1 | 1.73e-94 | 15 | 441 | 27 | 440 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4L4O_A | 3.77e-20 | 71 | 346 | 7 | 301 | Thecrystal structure of CbXyn10B in native form [Caldicellulosiruptor bescii DSM 6725] |
| 4PMD_A | 2.15e-19 | 71 | 346 | 7 | 301 | Crystalstructure of CbXyn10B from Caldicellulosiruptor bescii and its mutant(E139A) in complex with hydrolyzed xylotetraose [Caldicellulosiruptor bescii DSM 6725] |
| 4L4P_A | 2.33e-19 | 71 | 346 | 7 | 301 | themutant(E139A) structure in complex with xylotriose [Caldicellulosiruptor bescii DSM 6725] |
| 5Y3X_A | 2.18e-18 | 79 | 346 | 34 | 321 | Crystalstructure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_B Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_C Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_D Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_E Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_F Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL] |
| 6FHE_A | 3.15e-15 | 99 | 346 | 58 | 305 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
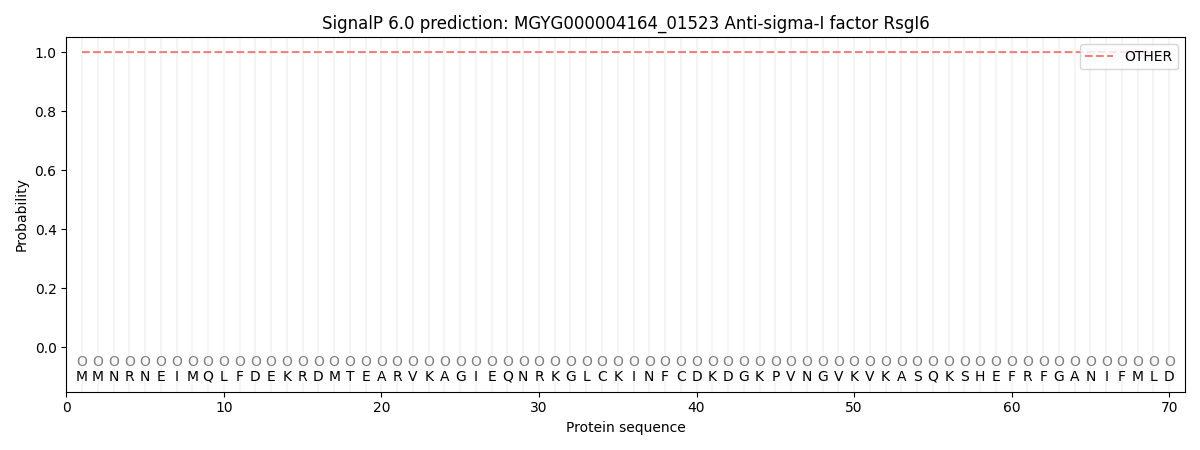
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000054 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
