You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003908_00765
You are here: Home > Sequence: MGYG000003908_00765
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Odoribacter sp900541415 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Marinifilaceae; Odoribacter; Odoribacter sp900541415 | |||||||||||
| CAZyme ID | MGYG000003908_00765 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | Alpha-xylosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 276087; End: 277667 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 139 | 523 | 4e-79 | 0.9110070257611241 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06592 | GH31_NET37 | 3.84e-171 | 134 | 496 | 1 | 364 | glucosidase NET37. NET37 (also known as KIAA1161) is a human lamina-associated nuclear envelope transmembrane protein. A member of the glycosyl hydrolase family 31 (GH31) , it has been shown to be required for myogenic differentiation of C2C12 cells. Related proteins are found in eukaryotes and prokaryotes. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| pfam01055 | Glyco_hydro_31 | 1.19e-54 | 148 | 522 | 40 | 435 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| COG1501 | YicI | 8.27e-52 | 57 | 524 | 187 | 665 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| PRK10426 | PRK10426 | 2.80e-31 | 182 | 522 | 260 | 616 | alpha-glucosidase; Provisional |
| cd06593 | GH31_xylosidase_YicI | 6.74e-31 | 138 | 298 | 11 | 172 | alpha-xylosidase YicI-like. YicI alpha-xylosidase is a glycosyl hydrolase family 31 (GH31) enzyme that catalyzes the release of an alpha-xylosyl residue from the non-reducing end of alpha-xyloside substrates such as alpha-xylosyl fluoride and isoprimeverose. YicI forms a homohexamer (a trimer of dimers). All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. The YicI family corresponds to subgroup 4 in the Ernst et al classification of GH31 enzymes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT50281.1 | 3.02e-254 | 3 | 526 | 8 | 531 |
| QCQ36424.1 | 2.52e-250 | 13 | 525 | 17 | 527 |
| QKH84612.1 | 1.02e-249 | 13 | 525 | 17 | 527 |
| QCQ51611.1 | 1.02e-249 | 13 | 525 | 17 | 527 |
| QCQ32141.1 | 1.02e-249 | 13 | 525 | 17 | 527 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2F2H_A | 9.59e-31 | 63 | 524 | 195 | 664 | Structureof the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_B Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_C Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_D Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_E Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_F Structure of the YicI thiosugar Michaelis complex [Escherichia coli] |
| 1XSI_A | 9.68e-31 | 63 | 524 | 195 | 664 | Structureof a Family 31 alpha glycosidase [Escherichia coli],1XSI_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_A Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSK_A Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_B Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_C Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_D Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_E Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_F Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli] |
| 1WE5_A | 3.28e-27 | 63 | 524 | 195 | 664 | CrystalStructure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_B Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_C Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_D Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_E Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_F Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli] |
| 5F7S_A | 5.57e-25 | 135 | 525 | 290 | 729 | Cycloalternan-degradingenzyme from Trueperella pyogenes [Trueperella pyogenes],5F7S_B Cycloalternan-degrading enzyme from Trueperella pyogenes [Trueperella pyogenes],5I0E_B Cycloalternan-degrading enzyme from Trueperella pyogenes in complex with isomaltose [Trueperella pyogenes],5I0F_B Cycloalternan-degrading enzyme from Trueperella pyogenes in complex with covalent intermediate [Trueperella pyogenes] |
| 5I0G_B | 5.54e-24 | 135 | 525 | 290 | 729 | Cycloalternan-degradingenzyme from Trueperella pyogenes in complex with cycloalternan [Trueperella pyogenes] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q6NSJ0 | 1.81e-57 | 103 | 522 | 283 | 710 | Myogenesis-regulating glycosidase OS=Homo sapiens OX=9606 GN=MYORG PE=1 SV=2 |
| Q69ZQ1 | 8.88e-56 | 103 | 522 | 282 | 712 | Myogenesis-regulating glycosidase OS=Mus musculus OX=10090 GN=Myorg PE=1 SV=2 |
| P31434 | 5.24e-30 | 63 | 524 | 195 | 664 | Alpha-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yicI PE=1 SV=2 |
| Q5AW25 | 1.79e-27 | 34 | 520 | 187 | 687 | Alpha-xylosidase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=agdD PE=1 SV=1 |
| P96793 | 1.79e-24 | 63 | 504 | 193 | 646 | Alpha-xylosidase XylQ OS=Lactiplantibacillus pentosus OX=1589 GN=xylQ PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
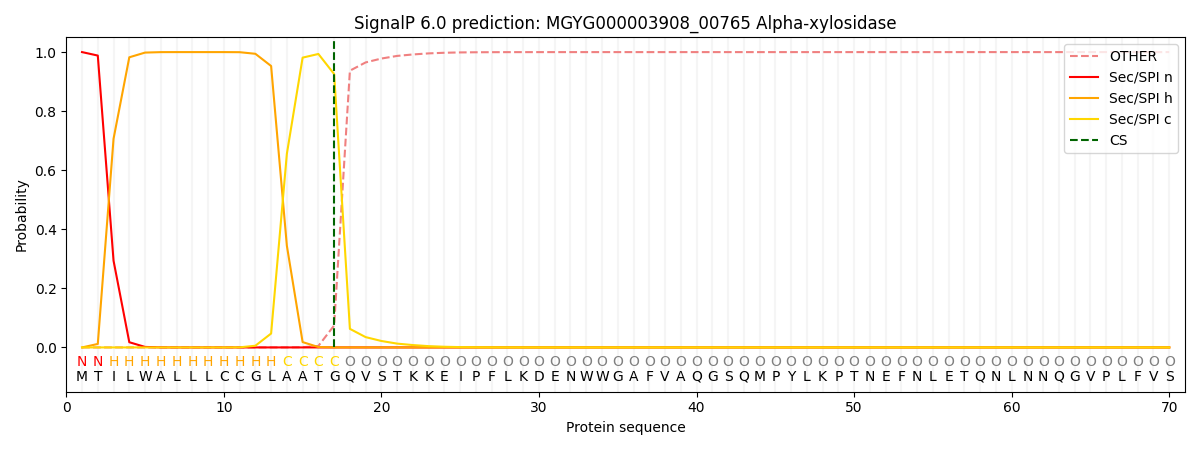
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000270 | 0.999165 | 0.000146 | 0.000141 | 0.000131 | 0.000128 |
