You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003835_02270
You are here: Home > Sequence: MGYG000003835_02270
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA7173 sp001915385 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; UBA7173; UBA7173 sp001915385 | |||||||||||
| CAZyme ID | MGYG000003835_02270 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Xylan 1,4-beta-xylosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7757; End: 10315 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 62 | 303 | 4.6e-66 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PLN03080 | PLN03080 | 1.21e-126 | 37 | 838 | 51 | 769 | Probable beta-xylosidase; Provisional |
| PRK15098 | PRK15098 | 3.82e-121 | 7 | 838 | 2 | 751 | beta-glucosidase BglX. |
| COG1472 | BglX | 6.35e-74 | 29 | 432 | 13 | 389 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 1.15e-48 | 75 | 336 | 73 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| pfam01915 | Glyco_hydro_3_C | 1.50e-48 | 376 | 737 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT73252.1 | 0.0 | 9 | 850 | 6 | 844 |
| BCA52477.1 | 0.0 | 9 | 850 | 6 | 844 |
| QUT77717.1 | 0.0 | 7 | 852 | 4 | 853 |
| ALJ40805.1 | 0.0 | 14 | 850 | 10 | 851 |
| AAO76885.1 | 0.0 | 14 | 850 | 10 | 851 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3AC0_A | 1.44e-85 | 40 | 845 | 7 | 829 | Crystalstructure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_B Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_C Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_D Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus] |
| 3ABZ_A | 1.29e-82 | 40 | 845 | 7 | 829 | Crystalstructure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_B Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_C Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_D Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus] |
| 5Z87_A | 2.17e-82 | 18 | 844 | 91 | 777 | ChainA, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
| 7VC7_A | 6.10e-79 | 32 | 822 | 19 | 703 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
| 7VC6_A | 6.10e-79 | 32 | 822 | 19 | 703 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EY15 | 4.42e-148 | 14 | 841 | 9 | 849 | Xylan 1,4-beta-xylosidase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyl3A PE=1 SV=1 |
| Q94KD8 | 1.17e-110 | 21 | 848 | 37 | 760 | Probable beta-D-xylosidase 2 OS=Arabidopsis thaliana OX=3702 GN=BXL2 PE=2 SV=1 |
| Q9LJN4 | 1.02e-104 | 22 | 842 | 35 | 762 | Probable beta-D-xylosidase 5 OS=Arabidopsis thaliana OX=3702 GN=BXL5 PE=2 SV=2 |
| Q9SGZ5 | 1.12e-101 | 38 | 811 | 48 | 730 | Probable beta-D-xylosidase 7 OS=Arabidopsis thaliana OX=3702 GN=BXL7 PE=2 SV=2 |
| Q9LXD6 | 9.47e-101 | 36 | 809 | 57 | 732 | Beta-D-xylosidase 3 OS=Arabidopsis thaliana OX=3702 GN=BXL3 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
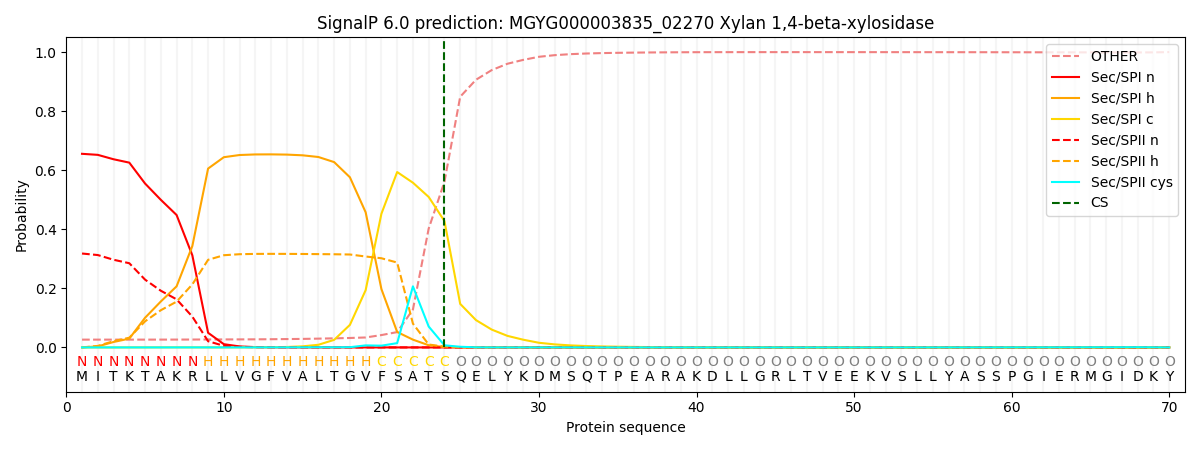
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.029222 | 0.647153 | 0.322802 | 0.000300 | 0.000249 | 0.000249 |
