You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003810_00563
You are here: Home > Sequence: MGYG000003810_00563
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
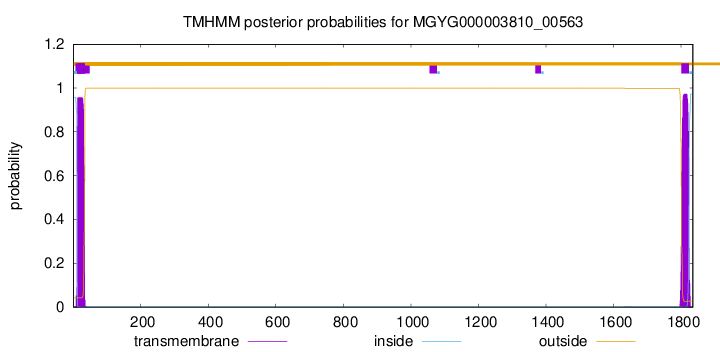
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Varibaculum; | |||||||||||
| CAZyme ID | MGYG000003810_00563 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 60245; End: 65752 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 47 | 412 | 1.5e-72 | 0.956140350877193 |
| GH20 | 908 | 1214 | 1.7e-51 | 0.9525222551928784 |
| CBM51 | 1464 | 1584 | 1.7e-32 | 0.8731343283582089 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 2.43e-84 | 45 | 416 | 2 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| cd06564 | GH20_DspB_LnbB-like | 1.08e-58 | 908 | 1214 | 2 | 326 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| pfam08305 | NPCBM | 3.09e-36 | 1443 | 1586 | 1 | 136 | NPCBM/NEW2 domain. This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. This domain has also been called the NEW2 domain (Naumoff DG. Phylogenetic analysis of alpha-galactosidases of the GH27 family. Molecular Biology (Engl Transl). (2004)38:388-399.) |
| smart00776 | NPCBM | 7.09e-29 | 1441 | 1586 | 1 | 145 | This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. |
| pfam00728 | Glyco_hydro_20 | 3.46e-28 | 922 | 1137 | 17 | 267 | Glycosyl hydrolase family 20, catalytic domain. This domain has a TIM barrel fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWG03116.1 | 5.11e-164 | 719 | 1771 | 256 | 1302 |
| AZR04729.1 | 5.11e-164 | 719 | 1771 | 256 | 1302 |
| AZR07627.1 | 5.11e-164 | 719 | 1771 | 256 | 1302 |
| AWG15845.1 | 5.11e-164 | 719 | 1771 | 256 | 1302 |
| QIU85951.1 | 1.54e-162 | 719 | 1771 | 250 | 1296 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4H04_A | 2.41e-132 | 764 | 1420 | 28 | 641 | Lacto-N-biosidasefrom Bifidobacterium bifidum [Bifidobacterium bifidum JCM 1254],4H04_B Lacto-N-biosidase from Bifidobacterium bifidum [Bifidobacterium bifidum JCM 1254],4JAW_A Crystal Structure of Lacto-N-Biosidase from Bifidobacterium bifidum complexed with LNB-thiazoline [Bifidobacterium bifidum JCM 1254],4JAW_B Crystal Structure of Lacto-N-Biosidase from Bifidobacterium bifidum complexed with LNB-thiazoline [Bifidobacterium bifidum JCM 1254],5BXP_A LNBase in complex with LNB-LOGNAc [Bifidobacterium bifidum JCM 1254],5BXP_B LNBase in complex with LNB-LOGNAc [Bifidobacterium bifidum JCM 1254],5BXR_A LNBase in complex with LNB-NHAcDNJ [Bifidobacterium bifidum JCM 1254],5BXR_B LNBase in complex with LNB-NHAcDNJ [Bifidobacterium bifidum JCM 1254],5BXS_A LNBase in complex with LNB-NHAcCAS [Bifidobacterium bifidum JCM 1254],5BXS_B LNBase in complex with LNB-NHAcCAS [Bifidobacterium bifidum JCM 1254],5BXT_A LNBase in complex with LNB-NHAcAUS [Bifidobacterium bifidum JCM 1254],5BXT_B LNBase in complex with LNB-NHAcAUS [Bifidobacterium bifidum JCM 1254] |
| 1EUT_A | 1.53e-86 | 50 | 509 | 18 | 444 | Sialidase,Large 68kd Form, Complexed With Galactose [Micromonospora viridifaciens],1EUU_A Sialidase Or Neuraminidase, Large 68kd Form [Micromonospora viridifaciens] |
| 1WCQ_A | 4.66e-86 | 50 | 509 | 14 | 440 | Mutagenesisof the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens],1WCQ_B Mutagenesis of the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens],1WCQ_C Mutagenesis of the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens] |
| 2BZD_A | 8.56e-86 | 50 | 509 | 14 | 440 | Galactoserecognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens],2BZD_B Galactose recognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens],2BZD_C Galactose recognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens] |
| 1W8N_A | 1.16e-85 | 50 | 509 | 14 | 440 | Contributionof the Active Site Aspartic Acid to Catalysis in the Bacterial Neuraminidase from Micromonospora viridifaciens. [Micromonospora viridifaciens],1W8O_A Contribution of the Active Site Aspartic Acid to Catalysis in the Bacterial Neuraminidase from Micromonospora viridifaciens [Micromonospora viridifaciens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02834 | 2.11e-86 | 3 | 509 | 9 | 486 | Sialidase OS=Micromonospora viridifaciens OX=1881 GN=nedA PE=1 SV=1 |
| B2UPR7 | 7.52e-24 | 844 | 1192 | 169 | 545 | Beta-hexosaminidase Amuc_2136 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_2136 PE=1 SV=1 |
| P49008 | 2.63e-12 | 783 | 1310 | 45 | 620 | Beta-hexosaminidase OS=Porphyromonas gingivalis (strain ATCC BAA-308 / W83) OX=242619 GN=nahA PE=3 SV=2 |
| P0C6E9 | 1.35e-11 | 211 | 389 | 549 | 750 | Sialidase OS=Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961) OX=243277 GN=nanH PE=1 SV=1 |
| A5F7A4 | 1.35e-11 | 211 | 389 | 549 | 750 | Sialidase OS=Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395) OX=345073 GN=nanH PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000726 | 0.997550 | 0.000243 | 0.000991 | 0.000272 | 0.000213 |