You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003282_00407
You are here: Home > Sequence: MGYG000003282_00407
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coprobacter sp900545915 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Coprobacteraceae; Coprobacter; Coprobacter sp900545915 | |||||||||||
| CAZyme ID | MGYG000003282_00407 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 106242; End: 108584 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 217 | 673 | 2e-138 | 0.9976580796252927 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01055 | Glyco_hydro_31 | 2.43e-165 | 218 | 673 | 1 | 442 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| cd06591 | GH31_xylosidase_XylS | 3.94e-157 | 237 | 566 | 1 | 322 | xylosidase XylS-like. XylS is a glycosyl hydrolase family 31 (GH31) alpha-xylosidase found in prokaryotes, eukaryotes, and archaea, that catalyzes the release of alpha-xylose from the non-reducing terminal side of the alpha-xyloside substrate. XylS has been characterized in Sulfolobus solfataricus where it hydrolyzes isoprimeverose, the p-nitrophenyl-beta derivative of isoprimeverose, and xyloglucan oligosaccharides, and has transxylosidic activity. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. The XylS family corresponds to subgroup 3 in the Ernst et al classification of GH31 enzymes. |
| COG1501 | YicI | 6.19e-157 | 25 | 742 | 55 | 740 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| cd06598 | GH31_transferase_CtsZ | 1.29e-73 | 237 | 575 | 1 | 328 | CtsZ (cyclic tetrasaccharide-synthesizing enzyme Z)-like. CtsZ is a bacterial 6-alpha-glucosyltransferase, first identified in Arthrobacter globiformis, that produces cyclic tetrasaccharides together with a closely related enzyme CtsY. CtsZ and CtsY both have a glycosyl hydrolase family 31 (GH31) catalytic domain; CtsY belongs to a different subfamily. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| cd06603 | GH31_GANC_GANAB_alpha | 5.16e-72 | 237 | 675 | 1 | 428 | neutral alpha-glucosidase C, neutral alpha-glucosidase AB. This subgroup includes the closely related glycosyl hydrolase family 31 (GH31) isozymes, neutral alpha-glucosidase C (GANC) and the alpha subunit of heterodimeric neutral alpha-glucosidase AB (GANAB). Initially distinguished on the basis of differences in electrophoretic mobility in starch gel, GANC and GANAB have been shown to have other differences, including those of substrate specificity. GANC and GANAB are key enzymes in glycogen metabolism that hydrolyze terminal, non-reducing 1,4-linked alpha-D-glucose residues from glycogen in the endoplasmic reticulum. The GANC/GANAB family includes the alpha-glucosidase II (ModA) from Dictyostelium discoideum as well as the alpha-glucosidase II (GLS2, or ROT2 - Reversal of TOR2 lethality protein 2) from Saccharomyces cerevisiae. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNL40761.1 | 0.0 | 27 | 779 | 31 | 785 |
| QDH54195.1 | 0.0 | 17 | 779 | 18 | 785 |
| QUT27035.1 | 0.0 | 27 | 779 | 31 | 785 |
| ALJ48502.1 | 0.0 | 27 | 779 | 31 | 785 |
| QRQ55338.1 | 0.0 | 27 | 779 | 31 | 785 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5JOU_A | 8.48e-153 | 196 | 765 | 355 | 936 | Bacteroidesovatus Xyloglucan PUL GH31 [Bacteroides ovatus],5JOV_A Bacteroides ovatus Xyloglucan PUL GH31 with bound 5FIdoF [Bacteroides ovatus] |
| 7KMP_A | 5.63e-147 | 36 | 754 | 80 | 946 | ChainA, Alpha-xylosidase [Xanthomonas citri pv. citri str. 306],7KNC_A Chain A, Alpha-xylosidase [Xanthomonas citri pv. citri str. 306] |
| 2XVG_A | 3.80e-133 | 36 | 777 | 69 | 984 | crystalstructure of alpha-xylosidase (GH31) from Cellvibrio japonicus [Cellvibrio japonicus],2XVK_A crystal structure of alpha-xylosidase (GH31) from Cellvibrio japonicus in complex with 5-fluoro-alpha-D-xylopyranosyl fluoride [Cellvibrio japonicus],2XVL_A crystal structure of alpha-xylosidase (GH31) from Cellvibrio japonicus in complex with Pentaerythritol propoxylate (5 4 PO OH) [Cellvibrio japonicus] |
| 6JR6_A | 2.78e-83 | 19 | 776 | 37 | 792 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR7_A Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101] |
| 6JR8_A | 3.75e-82 | 19 | 776 | 37 | 792 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9P999 | 2.75e-179 | 29 | 774 | 2 | 724 | Alpha-xylosidase OS=Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2) OX=273057 GN=xylS PE=1 SV=1 |
| A7LXT0 | 2.29e-152 | 8 | 765 | 11 | 935 | Alpha-xylosidase BoGH31A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02646 PE=1 SV=1 |
| Q9F234 | 3.98e-79 | 25 | 733 | 37 | 735 | Alpha-glucosidase 2 OS=Bacillus thermoamyloliquefaciens OX=1425 PE=3 SV=1 |
| A2QTU5 | 1.62e-65 | 140 | 669 | 159 | 725 | Alpha-xylosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=axlA PE=1 SV=1 |
| Q01336 | 2.49e-64 | 136 | 543 | 135 | 528 | Uncharacterized family 31 glucosidase ORF2 (Fragment) OS=Pseudescherichia vulneris OX=566 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
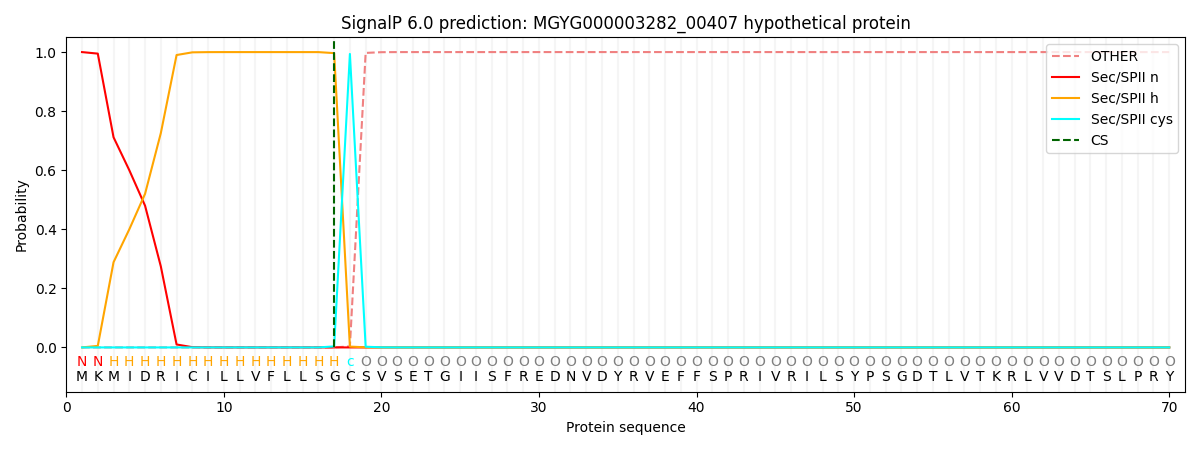
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000062 | 0.000000 | 0.000000 | 0.000000 |

