You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003154_01817
You are here: Home > Sequence: MGYG000003154_01817
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
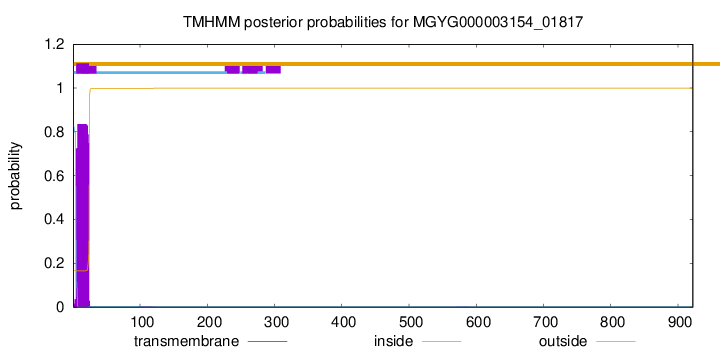
TMHMM annotations
Basic Information help
| Species | UMGS1490 sp900548185 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; UMGS1490; UMGS1490 sp900548185 | |||||||||||
| CAZyme ID | MGYG000003154_01817 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1704; End: 4472 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 54 | 596 | 7.5e-97 | 0.5877659574468085 |
| CBM42 | 816 | 920 | 3.1e-18 | 0.7573529411764706 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 1.07e-35 | 61 | 487 | 15 | 430 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 4.12e-26 | 50 | 460 | 3 | 417 | beta-D-glucuronidase; Provisional |
| PRK10340 | ebgA | 1.39e-20 | 125 | 486 | 113 | 472 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam00703 | Glyco_hydro_2 | 4.02e-16 | 231 | 322 | 2 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| pfam05270 | AbfB | 8.36e-16 | 822 | 920 | 39 | 137 | Alpha-L-arabinofuranosidase B (ABFB) domain. This family consists of several fungal alpha-L-arabinofuranosidase B proteins. L-Arabinose is a constituent of plant-cell-wall poly-saccharides. It is found in a polymeric form in L-arabinan, in which the backbone is formed by 1,5-a- linked l-arabinose residues that can be branched via 1,2-a- and 1,3-a-linked l-arabinofuranose side chains. AbfB hydrolyzes 1,5-a, 1,3-a and 1,2-a linkages in both oligosaccharides and polysaccharides, which contain terminal non-reducing l-arabinofuranoses in side chains. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATB39838.1 | 2.07e-199 | 13 | 624 | 21 | 604 |
| QRN98977.1 | 1.32e-194 | 23 | 626 | 33 | 607 |
| ADO72012.1 | 2.09e-192 | 2 | 626 | 12 | 605 |
| QRN98978.1 | 8.49e-192 | 23 | 624 | 33 | 603 |
| ATB43159.1 | 3.82e-190 | 23 | 626 | 33 | 607 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7SF2_A | 8.11e-160 | 24 | 624 | 5 | 587 | ChainA, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_E Chain E, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_F Chain F, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838] |
| 6S6Z_A | 7.31e-28 | 47 | 487 | 33 | 464 | Structureof beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_B Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_C Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_D Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_E Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_F Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_G Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_H Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8] |
| 6SD0_A | 7.32e-28 | 47 | 487 | 34 | 465 | Structureof beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_B Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_C Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_D Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8] |
| 6MVG_A | 2.76e-25 | 122 | 460 | 79 | 402 | Crystalstructure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus],6MVG_B Crystal structure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus],6MVG_C Crystal structure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus] |
| 3FN9_A | 8.91e-25 | 120 | 491 | 58 | 433 | Crystalstructure of putative beta-galactosidase from bacteroides fragilis [Bacteroides fragilis NCTC 9343],3FN9_B Crystal structure of putative beta-galactosidase from bacteroides fragilis [Bacteroides fragilis NCTC 9343],3FN9_C Crystal structure of putative beta-galactosidase from bacteroides fragilis [Bacteroides fragilis NCTC 9343],3FN9_D Crystal structure of putative beta-galactosidase from bacteroides fragilis [Bacteroides fragilis NCTC 9343] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KM09 | 3.21e-28 | 125 | 495 | 110 | 463 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22050 PE=2 SV=2 |
| Q56307 | 4.01e-27 | 47 | 487 | 34 | 465 | Beta-galactosidase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=lacZ PE=1 SV=2 |
| T2KPJ7 | 4.53e-26 | 56 | 484 | 53 | 464 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| P77989 | 1.19e-22 | 125 | 588 | 60 | 505 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
| P0C1Y0 | 7.37e-18 | 125 | 484 | 130 | 486 | Beta-galactosidase OS=Lactobacillus delbrueckii subsp. bulgaricus OX=1585 GN=lacZ PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
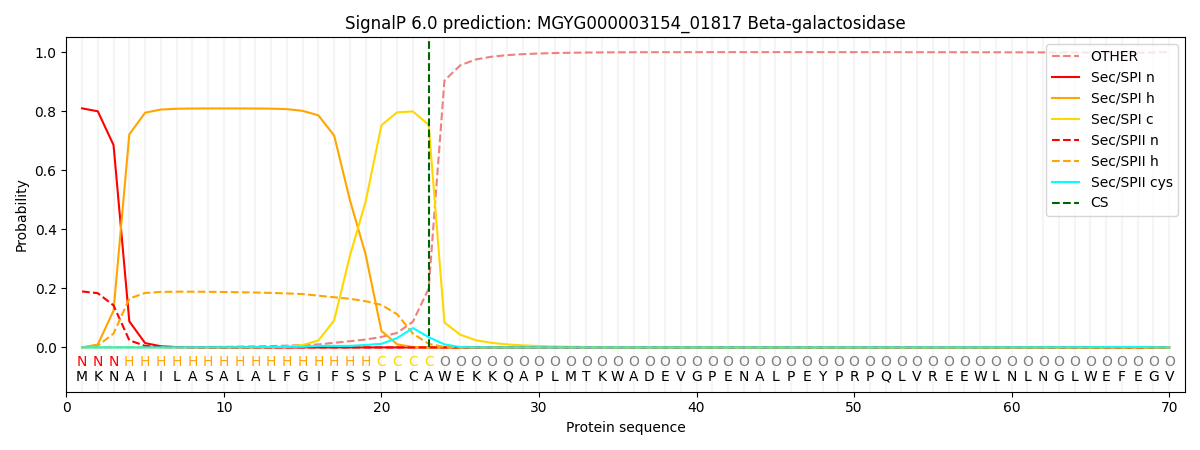
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.002533 | 0.799726 | 0.196754 | 0.000448 | 0.000261 | 0.000251 |