You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003145_00842
You are here: Home > Sequence: MGYG000003145_00842
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
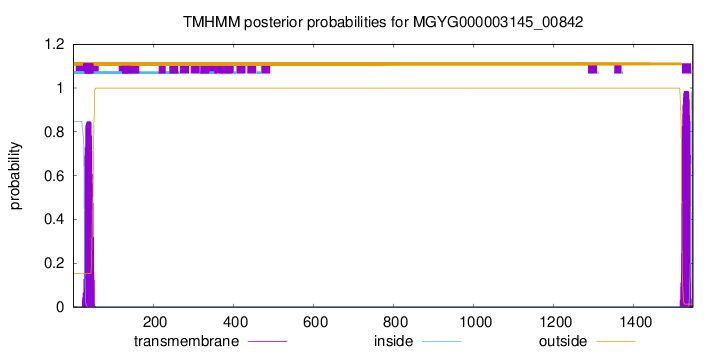
TMHMM annotations
Basic Information help
| Species | UMGS822 sp900545825 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; UMGS822; UMGS822 sp900545825 | |||||||||||
| CAZyme ID | MGYG000003145_00842 | |||||||||||
| CAZy Family | GH101 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 37886; End: 42532 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH101 | 245 | 899 | 1.2e-178 | 0.9943422913719944 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam12905 | Glyco_hydro_101 | 7.88e-139 | 485 | 757 | 1 | 273 | Endo-alpha-N-acetylgalactosaminidase. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae is largely determined by the ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. This family is the enzymatic region, EC:3.2.1.97, of the cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins. This reaction is exemplified by the S. pneumoniae protein Endo-alpha-N-acetylgalactosaminidase, where Asp764 is the catalytic nucleophile-base and Glu796 the catalytic proton donor. |
| cd14244 | GH_101_like | 6.84e-121 | 499 | 780 | 2 | 298 | Endo-a-N-acetylgalactosaminidase and related glcyosyl hydrolases. This family contains the enzymatically active domain of cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins (EC:3.2.1.97). It has been classified as glycosyl hydrolase family 101 in the Cazy resource. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae and other commensal human bacteria is largely determined by their ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. |
| pfam18080 | Gal_mutarotas_3 | 3.97e-38 | 241 | 484 | 1 | 243 | Galactose mutarotase-like fold domain. This domain is found in endo-alpha-N-acetylgalactosaminidase present in Streptococcus pneumoniae. Endo-alpha-N-acetylgalactosaminidase is a cell surface-anchored glycoside hydrolase involved in the breakdown of mucin type O-linked glycans. The domain, known as domain 2, exhibits strong structural similarlity to the galactose mutarotase-like fold but lacks the active site residues. Domains, found in a number of glycoside hydrolases, structurally similar to domain 2 confer stability to the multidomain architectures. |
| pfam17451 | Glyco_hyd_101C | 1.44e-37 | 763 | 872 | 1 | 111 | Glycosyl hydrolase 101 beta sandwich domain. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae is largely determined by the ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. This family is the enzymatic region, EC:3.2.1.97, of the cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins. This reaction is exemplified by a S. pneumoniae protein, where Asp764 is the catalytic nucleophile-base and Glu796 the catalytic proton donor. This domain represents C-terminal the beta sandwich domain. |
| pfam17974 | GalBD_like | 1.33e-22 | 1060 | 1256 | 1 | 184 | Galactose-binding domain-like. Proteins containing a galactose-binding domain-like fold can be found in several different protein families, in both eukaryotes and prokaryotes. The common function of these domains is to bind to specific ligands, such as cell-surface-attached carbohydrate substrates for galactose oxidase and sialidase, phospholipids on the outer side of the mammalian cell membrane for coagulation factor Va, membrane-anchored ephrin for the Eph family of receptor tyrosine kinases, and a complex of broken single-stranded DNA and DNA polymerase beta for XRCC1. The structure of the galactose-binding domain-like members consists of a beta-sandwich, in which the strands making up the sheets exhibit a jellyroll fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QXT63218.1 | 0.0 | 31 | 1263 | 3 | 1259 |
| AZR07626.1 | 0.0 | 19 | 1445 | 1 | 1566 |
| AWG15846.1 | 0.0 | 19 | 1445 | 1 | 1566 |
| AWG03117.1 | 0.0 | 19 | 1445 | 1 | 1566 |
| AZR04730.1 | 0.0 | 19 | 1445 | 1 | 1566 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2ZXQ_A | 1.39e-121 | 240 | 1430 | 26 | 1371 | Crystalstructure of endo-alpha-N-acetylgalactosaminidase from Bifidobacterium longum (EngBF) [Bifidobacterium longum] |
| 6QEP_A | 5.37e-115 | 240 | 1260 | 11 | 1114 | EngBFDARPin Fusion 4b H14 [Bifidobacterium longum] |
| 6QEV_B | 5.60e-115 | 240 | 1260 | 11 | 1114 | EngBFDARPin Fusion 4b B6 [Bifidobacterium longum] |
| 6QFK_A | 5.60e-115 | 240 | 1260 | 11 | 1114 | EngBFDARPin Fusion 4b G10 [Bifidobacterium longum] |
| 6SH9_B | 5.60e-115 | 240 | 1260 | 11 | 1114 | EngBFDARPin Fusion 4b D12 [Bifidobacterium longum subsp. longum JCM 1217] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A9WNA0 | 7.52e-199 | 239 | 1269 | 50 | 1044 | Putative endo-alpha-N-acetylgalactosaminidase OS=Renibacterium salmoninarum (strain ATCC 33209 / DSM 20767 / JCM 11484 / NBRC 15589 / NCIMB 2235) OX=288705 GN=RSal33209_1326 PE=3 SV=2 |
| Q2MGH6 | 3.18e-106 | 200 | 1372 | 290 | 1567 | Endo-alpha-N-acetylgalactosaminidase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=SP_0368 PE=1 SV=1 |
| Q8DR60 | 5.89e-105 | 200 | 1372 | 290 | 1567 | Endo-alpha-N-acetylgalactosaminidase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=spr0328 PE=1 SV=1 |
| Q02834 | 1.94e-18 | 1284 | 1430 | 504 | 643 | Sialidase OS=Micromonospora viridifaciens OX=1881 GN=nedA PE=1 SV=1 |
| I1S2N3 | 1.79e-09 | 1312 | 1430 | 71 | 186 | Galactose oxidase OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=GAOA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.067226 | 0.810040 | 0.002490 | 0.113972 | 0.005878 | 0.000396 |