You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002477_02172
You are here: Home > Sequence: MGYG000002477_02172
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Citrobacter_A amalonaticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Citrobacter_A; Citrobacter_A amalonaticus | |||||||||||
| CAZyme ID | MGYG000002477_02172 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2164831; End: 2165487 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16892 | LT_VirB1-like | 9.43e-69 | 9 | 153 | 3 | 142 | VirB1-like subfamily. This subfamily includes VirB1 protein, one of twelve proteins making up type IV secretion systems (T4SS). T4SS are macromolecular assemblies generally composed of VirB1-11 and VirD4 proteins, and are used by bacteria to transport material across their membranes. VirB1 acts as a lytic transglycosylase (LT), and is important with respect to piercing the peptidoglycan layer in the periplasm. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| PRK13864 | PRK13864 | 1.83e-13 | 10 | 181 | 36 | 194 | type IV secretion system lytic transglycosylase VirB1; Provisional |
| pfam01464 | SLT | 1.24e-04 | 23 | 108 | 18 | 80 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd13400 | LT_IagB-like | 3.38e-04 | 23 | 108 | 11 | 77 | Escherichia coli invasion protein IagB and similar proteins. Lytic transglycosylase-like protein, similar to Escherichia coli invasion protein IagB. IagB is encoded within a pathogenicity island in Salmonella enterica and has been shown to degrade polymeric peptidoglycan. IagB-like invasion proteins are implicated in the invasion of eukaryotic host cells by bacteria. Lytic transglycosylase (LT) catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Members of this family resemble the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| cd16893 | LT_MltC_MltE | 0.008 | 23 | 76 | 20 | 89 | membrane-bound lytic murein transglycosylases MltC and MltE, and similar proteins. MltC and MltE are periplasmic, outer membrane attached lytic transglycosylases (LTs), which cleave beta-1,4-glycosidic bonds joining N-acetylmuramic acid and N-acetylglucosamine in the cell wall peptidoglycan, yielding 1,6-anhydromuropeptides. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPB30960.1 | 2.61e-155 | 1 | 218 | 1 | 218 |
| AMG53476.1 | 2.61e-155 | 1 | 218 | 1 | 218 |
| BCU49487.1 | 2.61e-155 | 1 | 218 | 1 | 218 |
| QDK86790.1 | 2.61e-155 | 1 | 218 | 1 | 218 |
| AUZ67296.1 | 4.33e-154 | 1 | 218 | 1 | 218 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0C398 | 7.69e-45 | 1 | 149 | 1 | 150 | Type IV secretion system protein virB1 OS=Brucella abortus biovar 1 (strain 9-941) OX=262698 GN=virB1 PE=3 SV=1 |
| Q2YIT5 | 7.69e-45 | 1 | 149 | 1 | 150 | Type IV secretion system protein virB1 OS=Brucella abortus (strain 2308) OX=359391 GN=virB1 PE=3 SV=1 |
| Q8YDZ5 | 7.69e-45 | 1 | 149 | 1 | 150 | Type IV secretion system protein virB1 OS=Brucella melitensis biotype 1 (strain 16M / ATCC 23456 / NCTC 10094) OX=224914 GN=virB1 PE=3 SV=1 |
| Q9RPY4 | 7.69e-45 | 1 | 149 | 1 | 150 | Type IV secretion system protein virB1 OS=Brucella suis biovar 1 (strain 1330) OX=204722 GN=virB1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
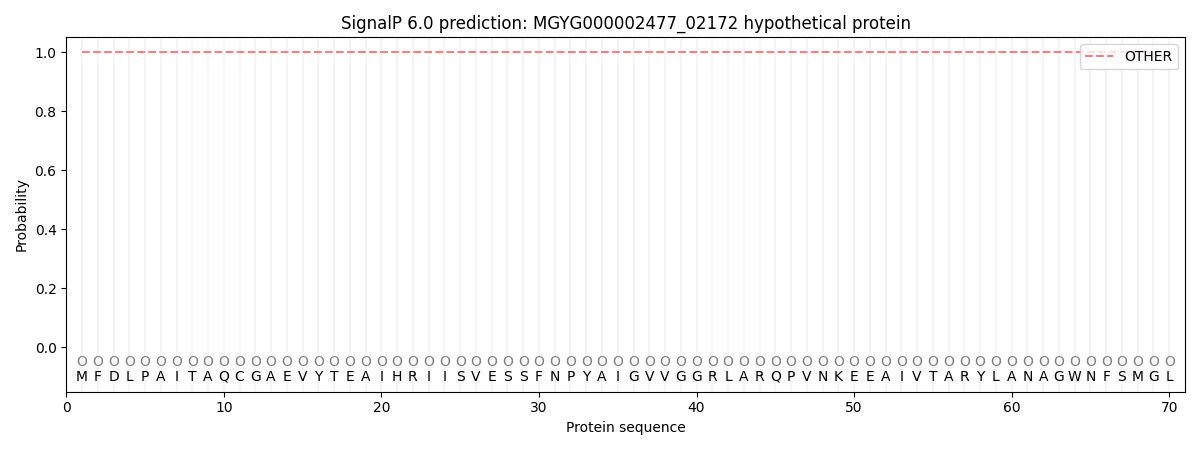
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000033 | 0.000015 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
