You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002270_03344
You are here: Home > Sequence: MGYG000002270_03344
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterococcus_A avium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus_A; Enterococcus_A avium | |||||||||||
| CAZyme ID | MGYG000002270_03344 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8570; End: 9733 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 32 | 357 | 8.2e-38 | 0.9239766081871345 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam13088 | BNR_2 | 3.36e-63 | 48 | 348 | 1 | 280 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| cd15482 | Sialidase_non-viral | 2.17e-60 | 31 | 364 | 8 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| COG4692 | COG4692 | 6.58e-37 | 36 | 363 | 32 | 371 | Predicted neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| COG4409 | NanH | 7.79e-07 | 134 | 368 | 495 | 726 | Neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| pfam15892 | BNR_4 | 6.17e-06 | 96 | 214 | 96 | 203 | BNR repeat-containing family member. BNR_4 is a family which carries the unique sequence motif SxDxGxTW which is so characteristic of the repeats of the BNR family, pfam02012. It is unclear whether or not this unit is repeated throughout the sequences of this family, but if it is then the family is likely to be bacterial neuraminidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCQ11920.1 | 2.27e-293 | 1 | 387 | 1 | 387 |
| QXJ61186.1 | 2.11e-271 | 3 | 387 | 4 | 388 |
| VEI59842.1 | 2.22e-246 | 1 | 387 | 2 | 388 |
| QEY07394.1 | 2.22e-246 | 1 | 387 | 2 | 388 |
| AWN18553.1 | 2.90e-244 | 1 | 387 | 1 | 387 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4FJ6_A | 5.59e-07 | 4 | 321 | 136 | 486 | Crystalstructure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_B Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_C Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_D Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
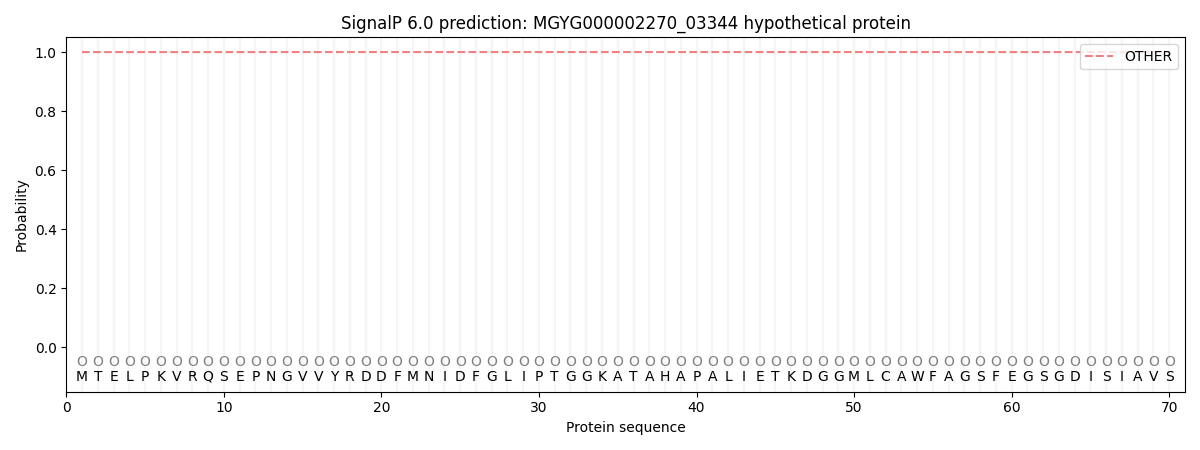
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000067 | 0.000007 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
