You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001949_00992
You are here: Home > Sequence: MGYG000001949_00992
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Dysosmobacter sp900551885 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; Dysosmobacter; Dysosmobacter sp900551885 | |||||||||||
| CAZyme ID | MGYG000001949_00992 | |||||||||||
| CAZy Family | SLH | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9204; End: 10892 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH25 | 352 | 545 | 4.1e-35 | 0.9943502824858758 |
| SLH | 27 | 66 | 4.6e-16 | 0.9285714285714286 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06414 | GH25_LytC-like | 1.49e-62 | 349 | 559 | 1 | 191 | The LytC lysozyme of Streptococcus pneumoniae is a bacterial cell wall hydrolase that cleaves the beta1-4-glycosydic bond located between the N-acetylmuramoyl-N-glucosaminyl residues of the cell wall polysaccharide chains. LytC is composed of a C-terminal glycosyl hydrolase family 25 (GH25) domain and an N-terminal choline-binding module (CBM) consisting of eleven homologous repeats that specifically recognizes the choline residues of pneumococcal lipoteichoic and teichoic acids. This domain arrangement is the reverse of the major pneumococcal autolysin, LytA, and the CPL-1-like lytic enzymes of the pneumococcal bacteriophages, in which the CBM (consisting of six repeats) is at the C-terminus. This model represents the C-terminal catalytic domain of the LytC-like enzymes. |
| cd00599 | GH25_muramidase | 2.12e-21 | 351 | 557 | 2 | 185 | Endo-N-acetylmuramidases (muramidases) are lysozymes (also referred to as peptidoglycan hydrolases) that degrade bacterial cell walls by catalyzing the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues. This family of muramidases contains a glycosyl hydrolase family 25 (GH25) catalytic domain and is found in bacteria, fungi, slime molds, round worms, protozoans and bacteriophages. The bacteriophage members are referred to as endolysins which are involved in lysing the host cell at the end of the replication cycle to allow release of mature phage particles. Endolysins are typically modular enzymes consisting of a catalytically active domain that hydrolyzes the peptidoglycan cell wall and a cell wall-binding domain that anchors the protein to the cell wall. Endolysins generally have narrow substrate specificities with either intra-species or intra-genus bacteriolytic activity. |
| pfam01183 | Glyco_hydro_25 | 7.39e-18 | 352 | 545 | 1 | 180 | Glycosyl hydrolases family 25. |
| NF033190 | inl_like_NEAT_1 | 2.13e-12 | 27 | 195 | 582 | 752 | NEAT domain-containing leucine-rich repeat protein. Members of this family have an N-terminal NEAT (near transporter) domain often associated with iron transport, followed by a leucine-rich repeat region with significant sequence similarity to the internalins of Listeria monocytogenes. However, since Bacillus cereus (from which this protein was described, in PMID:16978259) is not considered an intracellular pathogen, and the function may be iron transport rather than internalization, applying the name "internalin" to this family probably would be misleading. |
| cd06415 | GH25_Cpl1-like | 1.26e-11 | 351 | 557 | 3 | 193 | Cpl-1 lysin (also known as Cpl-9 lysozyme / muramidase) is a bacterial cell wall endolysin encoded by the pneumococcal bacteriophage Cp-1, which cleaves the glycosidic N-acetylmuramoyl-(beta1,4)-N-acetylglucosamine bonds of the pneumococcal glycan chain, thus acting as an enzymatic antimicrobial agent (an enzybiotic) against streptococcal infections. Cpl-1 belongs to the CP family of lysozymes (CPL lysozymes) which includes the Cpl-7 lysin. Cpl-1 has a glycosyl hydrolase family 25 (GH25) catalytic domain with an irregular (beta/alpha)5-beta3 barrel and a C-terminal cell wall-anchoring module formed by six similar choline-binding repeats (ChBr's). The ChBr's facilitate the anchoring of Cpl-1 to the choline-containing teichoic acid of the pneumococcal cell wall. Other members of this domain family have an N-terminal CHAP (cysteine, histidine-dependent amidohydrolases/peptidases) domain similar to that of the firmicute CHAP lysins and associated with endopeptidase activity. The Cpl-7 lysin is also included here as is LysB of Lactococcus phage, and the Mur lysin of Lactobacillus phage. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUO37824.1 | 0.0 | 1 | 559 | 1 | 556 |
| QCI60549.1 | 6.28e-281 | 1 | 562 | 1 | 546 |
| BCK85948.1 | 3.92e-271 | 1 | 562 | 1 | 550 |
| BAK99312.1 | 2.28e-208 | 1 | 560 | 1 | 542 |
| QNL45744.1 | 2.72e-197 | 15 | 562 | 14 | 545 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| C6CRV0 | 5.52e-10 | 34 | 191 | 1291 | 1455 | Endo-1,4-beta-xylanase A OS=Paenibacillus sp. (strain JDR-2) OX=324057 GN=xynA1 PE=1 SV=1 |
| Q06852 | 2.76e-08 | 49 | 195 | 2104 | 2264 | Cell surface glycoprotein 1 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=olpB PE=3 SV=2 |
| P19424 | 2.75e-07 | 40 | 195 | 56 | 214 | Endoglucanase OS=Bacillus sp. (strain KSM-635) OX=1415 PE=1 SV=1 |
| Q06853 | 7.39e-07 | 27 | 95 | 590 | 658 | Cell surface glycoprotein 2 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=Cthe_3079 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
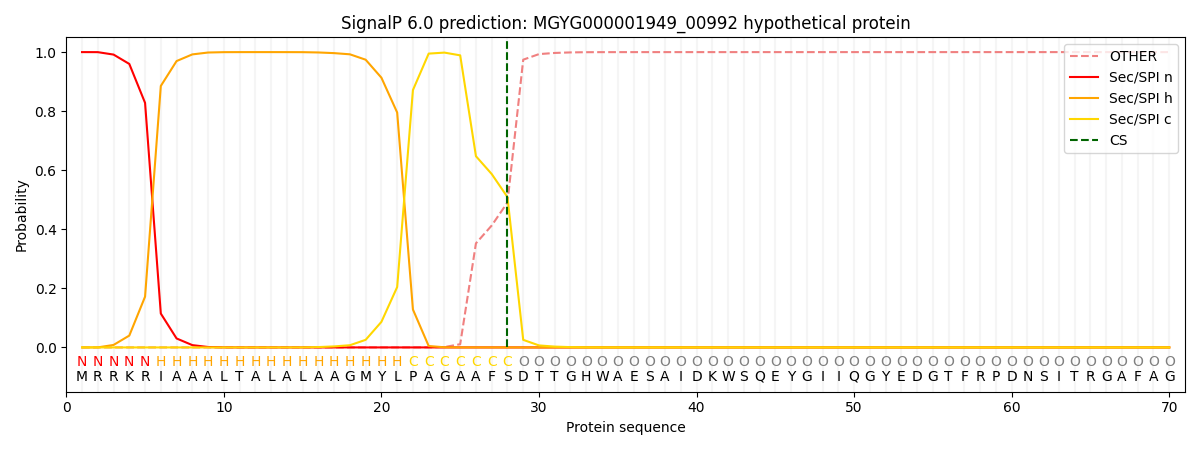
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000386 | 0.998703 | 0.000236 | 0.000259 | 0.000212 | 0.000190 |