You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001640_01692
You are here: Home > Sequence: MGYG000001640_01692
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Duncaniella sp900549445 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; Duncaniella; Duncaniella sp900549445 | |||||||||||
| CAZyme ID | MGYG000001640_01692 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8013; End: 9980 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3405 | BcsZ | 3.11e-44 | 192 | 576 | 1 | 353 | Endo-1,4-beta-D-glucanase Y [Carbohydrate transport and metabolism]. |
| pfam01270 | Glyco_hydro_8 | 3.31e-17 | 250 | 472 | 29 | 233 | Glycosyl hydrolases family 8. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUL54973.1 | 4.65e-134 | 145 | 601 | 123 | 564 |
| ASB49713.1 | 3.55e-125 | 187 | 654 | 298 | 769 |
| ADL50930.1 | 6.07e-122 | 192 | 574 | 32 | 406 |
| QNU67295.1 | 6.07e-122 | 192 | 575 | 31 | 407 |
| BAV13036.1 | 8.88e-122 | 192 | 574 | 44 | 418 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6G00_A | 2.28e-100 | 189 | 574 | 2 | 394 | CrystalStructure of a GH8 xylanase from Teredinibacter turnerae [Teredinibacter turnerae T7901],6G09_A Crystal Structure of a GH8 xylobiose complex from Teredinibacter turnerae [Teredinibacter turnerae T7901],6G0B_A Crystal Structure of a GH8 xylotriose complex from Teredinibacter Turnerae [Teredinibacter turnerae T7901] |
| 5YXT_A | 5.25e-100 | 191 | 574 | 5 | 376 | Glycosidehydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_B Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_C Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_D Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22] |
| 6G0N_A | 1.26e-99 | 189 | 574 | 2 | 394 | CrystalStructure of a GH8 catalytic mutant xylohexaose complex xylanase from Teredinibacter turnerae [Teredinibacter turnerae T7901] |
| 6SUD_A | 1.25e-97 | 191 | 574 | 6 | 380 | Structureof L320A mutant of Rex8A from Paenibacillus barcinonensis complexed with xylose. [Paenibacillus barcinonensis],6SUD_B Structure of L320A mutant of Rex8A from Paenibacillus barcinonensis complexed with xylose. [Paenibacillus barcinonensis] |
| 6SRD_A | 2.48e-97 | 191 | 574 | 6 | 380 | Structureof Rex8A from Paenibacillus barcinonensis complexed with xylose. [Paenibacillus barcinonensis],6SRD_B Structure of Rex8A from Paenibacillus barcinonensis complexed with xylose. [Paenibacillus barcinonensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A0S2UQQ5 | 1.28e-96 | 191 | 574 | 6 | 380 | Reducing-end xylose-releasing exo-oligoxylanase Rex8A OS=Paenibacillus barcinonensis OX=198119 GN=rex8A PE=1 SV=1 |
| Q9KB30 | 1.27e-87 | 192 | 574 | 7 | 379 | Reducing end xylose-releasing exo-oligoxylanase OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=BH2105 PE=1 SV=1 |
| A1A048 | 3.29e-74 | 201 | 572 | 14 | 377 | Reducing end xylose-releasing exo-oligoxylanase OS=Bifidobacterium adolescentis (strain ATCC 15703 / DSM 20083 / NCTC 11814 / E194a) OX=367928 GN=xylA PE=1 SV=1 |
| P37701 | 1.70e-26 | 250 | 571 | 93 | 389 | Endoglucanase 2 OS=Ruminiclostridium josui OX=1499 GN=celB PE=3 SV=1 |
| P37699 | 1.05e-24 | 250 | 571 | 93 | 389 | Endoglucanase C OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCC PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
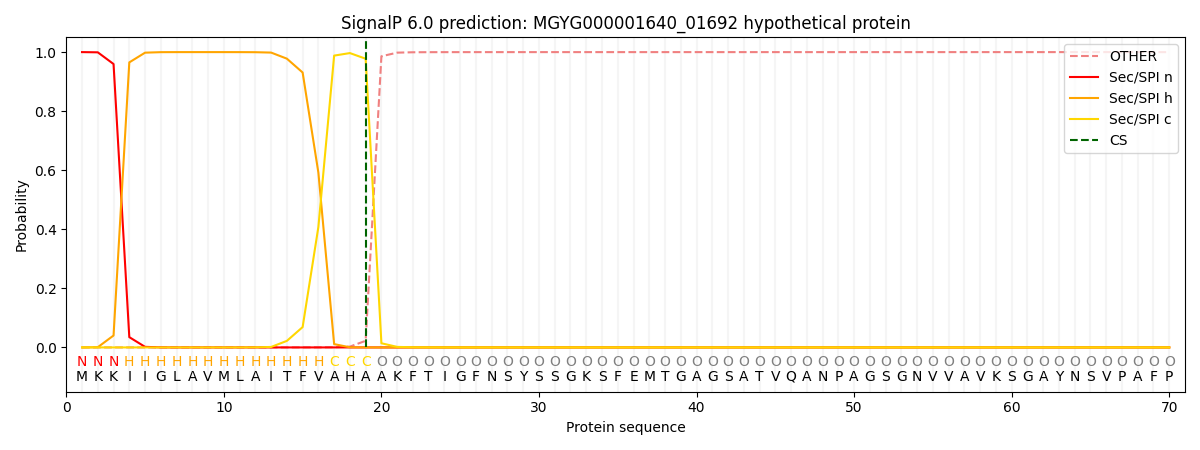
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000336 | 0.998834 | 0.000257 | 0.000187 | 0.000181 | 0.000172 |