You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001508_02772
You are here: Home > Sequence: MGYG000001508_02772
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
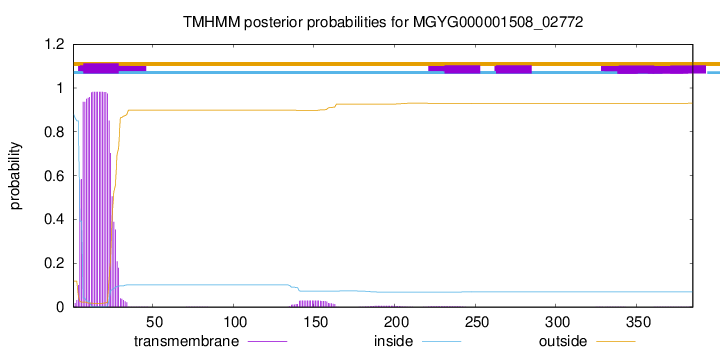
TMHMM annotations
Basic Information help
| Species | Thalassobacillus devorans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_D; Halobacillaceae; Thalassobacillus; Thalassobacillus devorans | |||||||||||
| CAZyme ID | MGYG000001508_02772 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1940984; End: 1942141 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 36 | 159 | 6.5e-32 | 0.9230769230769231 |
| CBM50 | 292 | 335 | 3.9e-18 | 0.975 |
| CBM50 | 341 | 384 | 1.1e-17 | 0.975 |
| CBM50 | 239 | 282 | 3.3e-16 | 0.975 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10917 | CE4_NodB_like_6s_7s | 2.03e-72 | 41 | 215 | 1 | 171 | Catalytic NodB homology domain of rhizobial NodB-like proteins. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes many rhizobial NodB chitooligosaccharide N-deacetylase (EC 3.5.1.-)-like proteins, mainly from bacteria and eukaryotes, such as chitin deacetylases (EC 3.5.1.41), bacterial peptidoglycan N-acetylglucosamine deacetylases (EC 3.5.1.-), and acetylxylan esterases (EC 3.1.1.72), which catalyze the N- or O-deacetylation of substrates such as acetylated chitin, peptidoglycan, and acetylated xylan. All members of this family contain a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold with 6- or 7 strands. Their catalytic activity is dependent on the presence of a divalent cation, preferably cobalt or zinc, and they employ a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. Several family members show diversity both in metal ion specificities and in the residues that coordinate the metal. |
| TIGR02764 | spore_ybaN_pdaB | 3.00e-68 | 36 | 228 | 1 | 191 | polysaccharide deacetylase family sporulation protein PdaB. This model describes the YbaN protein family, also called PdaB and SpoVIE, of Gram-positive bacteria. Although ybaN null mutants have only a mild sporulation defect, ybaN/ytrI double mutants show drastically reducted sporulation efficiencies. This synthetic defect suggests the role of this sigmaE-controlled gene in sporulation had been masked by functional redundancy. Members of this family are homologous to a characterized polysaccharide deacetylase; the exact function this protein family is unknown. [Cellular processes, Sporulation and germination] |
| cd10954 | CE4_CtAXE_like | 9.88e-59 | 41 | 226 | 1 | 179 | Catalytic NodB homology domain of Clostridium thermocellum acetylxylan esterase and its bacterial homologs. This family is represented by Clostridium thermocellum acetylxylan esterase (CtAXE, EC 3.1.1.72), a member of the carbohydrate esterase 4 (CE4) superfamily. CtAXE deacetylates O-acetylated xylan, a key component of plant cell walls. It shows no detectable activity on generic esterase substrates including para-nitrophenyl acetate. It is specific for sugar-based substrates and will precipitate acetylxylan, as a consequence of deacetylation. CtAXE is a monomeric protein containing a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold as other CE4 esterases. However, due to differences in the topography of the substrate-binding groove, the chemistry of the active center, and metal ion coordination, CtAXE has different metal ion preference and lacks activity on N-acetyl substrates. It is significantly activated by Co2+. Moreover, CtAXE displays distinctly different ligand coordination to the metal ion, utilizing an aspartate, a histidine, and four water molecules, as opposed to the conserved His-His-Asp zinc-binding triad of other CE4 esterases. |
| cd10950 | CE4_BsYlxY_like | 1.23e-57 | 37 | 228 | 2 | 188 | Putative catalytic NodB homology domain of uncharacterized protein YlxY from Bacillus subtilis and its bacterial homologs. The Bacillus subtilis genome contains six polysaccharide deacetylase gene homologs: pdaA, pdaB (previously known as ybaN), yheN, yjeA, yxkH and ylxY. This family is represented by Bacillus subtilis putative polysaccharide deacetylase BsYlxY, encoded by the ylxY gene, which is a member of the carbohydrate esterase 4 (CE4) superfamily. Although its biological function still remains unknown, BsYlxY shows high sequence homology to the catalytic domain of Bacillus subtilis pdaB gene encoding a putative polysaccharide deacetylase (BsPdaB), which is essential for the maintenance of spores after the late stage of sporulation and is highly conserved in spore-forming bacteria. However, disruption of the ylxY gene in B. subtilis did not cause any sporulation defect. Moreover, the Asp residue in the classical His-His-Asp zinc-binding motif of CE4 esterases is mutated to a Val residue in this family. Other catalytically relevant residues of CE4 esterases are also not conserved, which suggest that members of this family may be inactive. |
| COG0726 | CDA1 | 2.12e-54 | 19 | 228 | 45 | 257 | Peptidoglycan/xylan/chitin deacetylase, PgdA/CDA1 family [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCG43929.1 | 2.58e-150 | 13 | 335 | 14 | 339 |
| ASF38113.1 | 2.58e-150 | 13 | 335 | 14 | 339 |
| ANU13868.1 | 1.30e-140 | 2 | 340 | 4 | 346 |
| ANU22587.1 | 2.45e-138 | 20 | 338 | 22 | 344 |
| ANU10225.1 | 7.28e-136 | 1 | 335 | 2 | 339 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7FBW_A | 1.19e-33 | 38 | 228 | 114 | 300 | ChainA, Predicted xylanase/chitin deacetylase [Caldanaerobacter subterraneus subsp. tengcongensis MB4] |
| 5O6Y_A | 1.38e-33 | 43 | 224 | 23 | 204 | Crystalstructure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579] |
| 5O6Y_B | 1.99e-32 | 43 | 224 | 23 | 204 | Crystalstructure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579],5O6Y_C Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579],5O6Y_D Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579] |
| 4L1G_A | 8.44e-32 | 43 | 224 | 75 | 256 | Crystalstructure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase from Bacillus cereus [Bacillus cereus ATCC 14579],4L1G_B Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase from Bacillus cereus [Bacillus cereus ATCC 14579],4L1G_C Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase from Bacillus cereus [Bacillus cereus ATCC 14579],4L1G_D Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase from Bacillus cereus [Bacillus cereus ATCC 14579] |
| 1W1A_1 | 2.18e-31 | 37 | 228 | 56 | 247 | Structureof Bacillus subtilis PdaA in complex with NAG, a family 4 Carbohydrate esterase. [Bacillus subtilis],1W1A_2 Structure of Bacillus subtilis PdaA in complex with NAG, a family 4 Carbohydrate esterase. [Bacillus subtilis],1W1B_1 Structure of Bacillus subtilis PdaA with Cadmium, a family 4 Carbohydrate esterase. [Bacillus subtilis],1W1B_2 Structure of Bacillus subtilis PdaA with Cadmium, a family 4 Carbohydrate esterase. [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q81EK9 | 3.44e-32 | 43 | 224 | 83 | 264 | Peptidoglycan-N-acetylglucosamine deacetylase BC_1960 OS=Bacillus cereus (strain ATCC 14579 / DSM 31 / CCUG 7414 / JCM 2152 / NBRC 15305 / NCIMB 9373 / NCTC 2599 / NRRL B-3711) OX=226900 GN=BC_1960 PE=1 SV=1 |
| O34928 | 1.38e-30 | 37 | 228 | 62 | 253 | Peptidoglycan-N-acetylmuramic acid deacetylase PdaA OS=Bacillus subtilis (strain 168) OX=224308 GN=pdaA PE=1 SV=1 |
| Q8DP63 | 2.15e-29 | 41 | 229 | 268 | 449 | Peptidoglycan-N-acetylglucosamine deacetylase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=pgdA PE=1 SV=1 |
| Q04729 | 5.33e-28 | 39 | 228 | 65 | 254 | Uncharacterized 30.6 kDa protein in fumA 3'region OS=Geobacillus stearothermophilus OX=1422 PE=3 SV=1 |
| P50850 | 1.15e-27 | 33 | 229 | 122 | 313 | Uncharacterized protein YlxY OS=Bacillus subtilis (strain 168) OX=224308 GN=ylxY PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
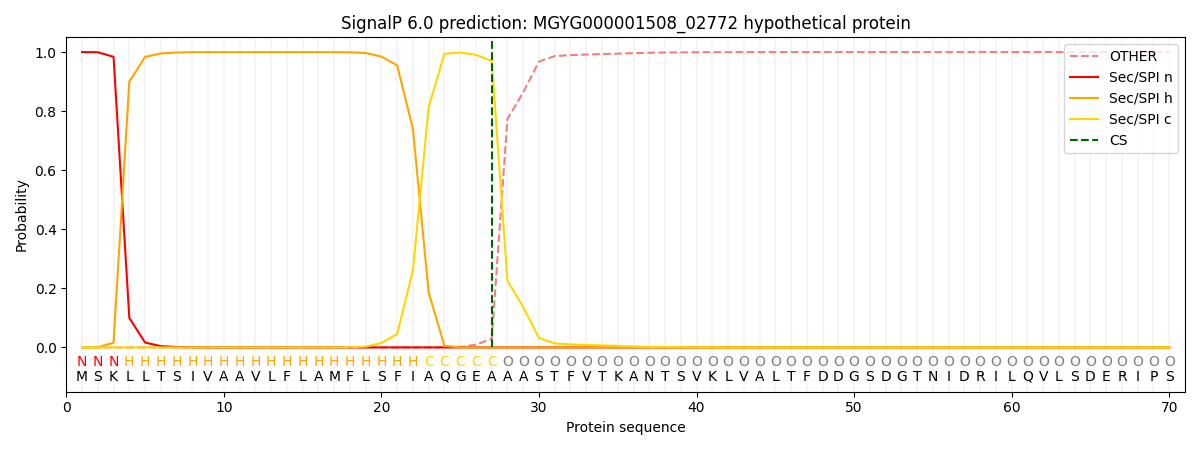
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000269 | 0.999109 | 0.000164 | 0.000154 | 0.000147 | 0.000139 |