You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001106_00584
You are here: Home > Sequence: MGYG000001106_00584
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
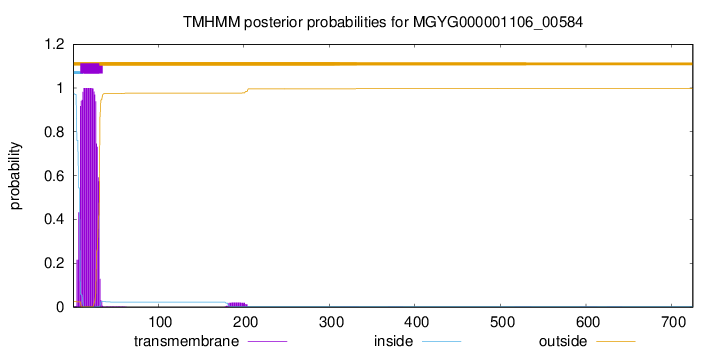
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Pygmaiobacter; | |||||||||||
| CAZyme ID | MGYG000001106_00584 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Penicillin-binding protein 1F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1463; End: 3640 Strand: - | |||||||||||
Full Sequence Download help
| MWANIGRVLV VLVSIGVIVA SVLAVMLSLY VVKVTANDAT LLSLENLKLS YSSTIYAINA | 60 |
| QTGEPEPYQR LVDNADRIWV DGNQIPDNLK NAFIAVEDKT FRENNGVNFK RTVSATANLV | 120 |
| LSKLTGGKIK LYDTEQGAST ITQQLIKNIT GDDDHDALRK VREIFRAISL TGRYSKDTIL | 180 |
| EAYLNTVGFM GNTCGVQACA YNLYDKDVSQ LTLAQCATVA AITNNPTKYN PRTEPENNLQ | 240 |
| RRNMILGMML EQGLITQAEH DAAVAEPLNL IDPAEKQNTA KVGNNSYFTD MVIDEVITDL | 300 |
| VSQGICETRS EASALLYNGG LAIYSTVDTN LQSIMEEVMA KEPGNYYPDQ EVEYEDKATG | 360 |
| ETKTQNVQGA MITIDYTGAV KGVVGGLGEK TADRGLNRAV DSLRQTGSTM KPIGAYAMGI | 420 |
| DYNYITYSDG GFIDDWVEEI EDKGSMRKWP VNYSGRYTEQ RMLVCDALAN SINTIAVRVG | 480 |
| KLVGPDDMFD FMTNTLGITS LVESRTEADG KISTDIGVGP LVLGSMTDGV SLAEMAHAYM | 540 |
| MFGDGGTVTP LHSYTTVENV QTGEVVLDNT LLPTTRAIGE DTAMIMNKLL RQVITRGTAA | 600 |
| GTGTNTIAGM ETVGKTGTTS NDKDHWFIGL TPYYVTATWM GYDQPQELAW KAYGKHPPTL | 660 |
| AWKEVMERAQ EGLEAKNFPT APAGTVVTQA YCTESGGAAG PLCPKTATGY YKADQLPEPC | 720 |
| DIHVG | 725 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 73 | 249 | 7.3e-54 | 0.943502824858757 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0744 | MrcB | 4.53e-149 | 5 | 701 | 11 | 628 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| TIGR02074 | PBP_1a_fam | 4.97e-142 | 76 | 672 | 1 | 531 | penicillin-binding protein, 1A family. Bacterial that synthesize a cell wall of peptidoglycan (murein) generally have several transglycosylases and transpeptidases for the task. This family consists of bifunctional transglycosylase/transpeptidase penicillin-binding proteins (PBP). In the Proteobacteria, this family includes PBP 1A but not the paralogous PBP 1B (TIGR02071). This family also includes related proteins, often designated PBP 1A, from other bacterial lineages. [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan] |
| COG5009 | MrcA | 1.30e-102 | 5 | 724 | 5 | 772 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
| TIGR02071 | PBP_1b | 1.42e-77 | 76 | 692 | 146 | 712 | penicillin-binding protein 1B. Bacterial that synthesize a cell wall of peptidoglycan (murein) generally have several transglycosylases and transpeptidases for the task. This family consists of a particular bifunctional transglycosylase/transpeptidase in E. coli and other Proteobacteria, designated penicillin-binding protein 1B. [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan] |
| pfam00912 | Transgly | 4.92e-60 | 77 | 250 | 13 | 177 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL02306.1 | 3.49e-172 | 5 | 720 | 46 | 796 |
| ATL89188.1 | 8.64e-172 | 6 | 721 | 37 | 787 |
| AXB28952.1 | 1.52e-171 | 5 | 720 | 61 | 811 |
| ATO98888.1 | 2.43e-171 | 6 | 721 | 37 | 787 |
| QIA44319.1 | 4.01e-171 | 6 | 721 | 54 | 804 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3UDF_A | 1.80e-46 | 78 | 675 | 40 | 721 | ChainA, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDF_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii] |
| 3ZG8_B | 6.44e-44 | 174 | 644 | 7 | 434 | CrystalStructure of Penicillin Binding Protein 4 from Listeria monocytogenes in the Ampicillin bound form [Listeria monocytogenes],3ZG9_B Crystal Structure of Penicillin-Binding Protein 4 from Listeria monocytogenes in the Cefuroxime bound form [Listeria monocytogenes],3ZGA_B Crystal Structure of Penicillin-Binding Protein 4 from Listeria monocytogenes in the Carbenicillin bound form [Listeria monocytogenes] |
| 5FGZ_A | 2.73e-43 | 77 | 654 | 156 | 678 | E.coli PBP1b in complex with FPI-1465 [Escherichia coli K-12],5HL9_A E. coli PBP1b in complex with acyl-ampicillin and moenomycin [Escherichia coli K-12],5HLA_A E. coli PBP1b in complex with acyl-cephalexin and moenomycin [Escherichia coli K-12],5HLB_A E. coli PBP1b in complex with acyl-aztreonam and moenomycin [Escherichia coli K-12],5HLD_A E. coli PBP1b in complex with acyl-CENTA and moenomycin [Escherichia coli K-12],6YN0_A Structure of E. coli PBP1b with a FtsN peptide activating transglycosylase activity [Escherichia coli K-12],7LQ6_A Chain A, Penicillin-binding protein 1B [Escherichia coli K-12] |
| 3VMA_A | 3.15e-43 | 77 | 654 | 177 | 699 | CrystalStructure of the Full-Length Transglycosylase PBP1b from Escherichia coli [Escherichia coli K-12] |
| 7U4H_A | 1.11e-41 | 69 | 678 | 32 | 769 | ChainA, Penicillin-binding protein 1A (Pbp1a) [Chlamydia trachomatis D/UW-3/CX],7U4H_B Chain B, Penicillin-binding protein 1A (Pbp1a) [Chlamydia trachomatis D/UW-3/CX] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0SRL7 | 3.45e-72 | 71 | 711 | 85 | 708 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain SM101 / Type A) OX=289380 GN=pbpA PE=3 SV=1 |
| Q0TNZ8 | 2.00e-71 | 71 | 711 | 85 | 708 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=pbpA PE=3 SV=1 |
| Q8XJ01 | 7.36e-71 | 71 | 648 | 85 | 646 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=pbpA PE=3 SV=1 |
| A7GHV1 | 2.41e-65 | 70 | 676 | 69 | 650 | Penicillin-binding protein 1A OS=Clostridium botulinum (strain Langeland / NCTC 10281 / Type F) OX=441772 GN=pbpA PE=3 SV=1 |
| Q891X1 | 2.75e-65 | 71 | 725 | 74 | 715 | Penicillin-binding protein 1A OS=Clostridium tetani (strain Massachusetts / E88) OX=212717 GN=pbpA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.957581 | 0.036620 | 0.000377 | 0.000330 | 0.000160 | 0.004964 |