You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001063_05442
You are here: Home > Sequence: MGYG000001063_05442
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Robinsoniella sp900540475 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Robinsoniella; Robinsoniella sp900540475 | |||||||||||
| CAZyme ID | MGYG000001063_05442 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 74926; End: 80409 Strand: + | |||||||||||
Full Sequence Download help
| MKRKNGFARG VSFMLSMALV LNTIVMPVSA SGQNGAAQVS AGGQVLEIEY EQAEKRGTET | 60 |
| EINQPVRDAA AAAQQADAEM ENNSDTGIDS DSGIGTDTGI DTETGTNFVT EAEIDSEKVT | 120 |
| EADTETELNG GQEIITEKEK DIDAATVTEK ESGLDQEPES VTGENTVKAL TDMEIGLFGE | 180 |
| IFNTEGYRGG SMELFNFENK TGEQVFASLS GQNLRPVFDK YNGNKGDEQF ATARFTGKLN | 240 |
| VEKAGDYKFF GWGDDGFRIW IDGKLVIDFW VQEWEKEQTS KAVALSEGLH DIKIEYLQGW | 300 |
| GGAVLKLSWE SANANIAKEV IPEKAFYHAK TPADTGLTAQ IYKVNGKPSS ITSFGFAGEP | 360 |
| VNQVLPNLDG HDGLKEAIQL SNGSSEYATA IIHGYITPQE SGDYEFVMSG DDAFRFWMKD | 420 |
| EMMVNSWKLH SDQTKVSQPV HMEKGFLYKI RINYAQADGD ANLNLSWKKD GGQAQVVPAE | 480 |
| CFRQAAKAIQ GETAAEDQHG LLAEVYGDIF LEEKKGEVIA PVLDKSLSIK KILSQYNGSE | 540 |
| EYGSFRFTGQ VNAEEAGDYT FYLIGDDGFR LWVNDELAID FWESKWDKQQ ESQPIHLNEG | 600 |
| RNDIRIEYFQ GFGGAYIMLE WLKPGAEEKE VIPTENLYPA STKKGLIGEI YQEKGGNQEK | 660 |
| LAEVLANEVS SEQFSVNSLV SYYEGNENAP ANVELNGKIK PEVSGTYEFA FGSSGNYQVY | 720 |
| LNDEKIISQN AGNVYKETQS AKVELEKGIY YNLTIIAENL NLKEDINLSW TRDGVKSVIG | 780 |
| EDQLYAPEAM WYESVTAREK LYVGLTEAVA LRRNTKVGSG EGQIAKEHMD KFSAVVDEIA | 840 |
| ADAKDFDIIA SVLKEDTNKI TDAVAEFKLN VIRVQKGEAL TQFNNGLYQG QDPFISYHDG | 900 |
| YYYFVSSSND PSHNKMYVSK SKSLLDQGEK VRIFDFNDSK SRIFAPEMFF IDGKWYIYYC | 960 |
| ADAKEYNWRH MACVMESVTD DPQGDWIDRG VLYTGMEIHN PNEFAQGNDF TVFQYRGQLY | 1020 |
| ACWGSMESGV ESPAIARMES PIKITEERSF LPGFGGEGPR AVVNGDHLFM TVSQGSFKTA | 1080 |
| GYHMGMYIFD TAKDDLLNPD NWSYKEGVFE GTKDVYGPAR ASFFKSADGT EDWMAFHSKV | 1140 |
| YPSNNNAWRQ VSIKKITWNE DGSPNFGRPV SPYEFTALPS GDPGLGLAYQ AEDGDLSGGA | 1200 |
| KKGYKGKGFQ GTGYGNITDT LGEKVTFNVN APADGDYFVR ARYANGVKAP ISATDNRGEW | 1260 |
| EANIPDITPK AGTLNLYVND AGQKSVTVSF DKTETDWQHW MTVCSRVTLK KGENTISYVK | 1320 |
| DAENSGNVYL DYITVQEAHE QPAAQTVKTG AELDALIAEV EGFRQEAYSI ETWRIFSAAL | 1380 |
| LSAKSVNKNA VQAVSNRYLS LEAAAKDLSN YVRFHADKNV SVSGDVKDDM YRIGSEVTVQ | 1440 |
| AKDLAGKTFS RWDVSGFELT EEQKVSKEFT FTVPRNRGSI VPVYAENEYY KVTAGTNVRI | 1500 |
| DNAAANSLYQ AGTTVKATVE IPQGKEFIGW DVKGITLTEA EKMAVTISFV MPESEVVLNA | 1560 |
| VFKNIVIPEK PKYQLKAGAG ISIANAAKDQ MYTAGTRIQA TVSVPAGKVF VQWKVSGISL | 1620 |
| TAQQQKSTVI SFLMPASSVS LEAVFDIQKK KQTITAKDFQ KTDGDKSFYI KAKASGKGKL | 1680 |
| SYKSDHTKVV KVDSKGKVTI KGTGKAQITI TAAGNSSYDS AIKKITISVA PKKTADVKAV | 1740 |
| SKKSKTLKLT WKKTRGASGY LISYSTDKKF KKGVKKVVIS SGKTTSRTIK NLKGKKTCYV | 1800 |
| KVRAFKTSGK SRVYGKYSKT VSVRVKK | 1827 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 883 | 1164 | 4.6e-61 | 0.9965986394557823 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd18820 | GH43_LbAraf43-like | 2.61e-74 | 891 | 1139 | 1 | 258 | Glycosyl hydrolase family 43 proteins similar to Lactobacillus brevis alpha-L-arabinofuranosidase LbAraf43 and Geobacillus thermoleovorans GbtXyl43B. This uncharacterized glycosyl hydrolase family 43 (GH43) subgroup belongs to a subgroup which includes enzymes with beta-xylosidase (EC 3.2.1.37), alpha-L-arabinofuranosidase (EC 3.2.1.55) and possibly bifunctional xylosidase/arabinofuranosidase activities, similar to Lactobacillus brevis alpha-L-arabinofuranosidase LbAraf43 and Geobacillus thermoleovorans IT-08 beta-xylosidase / exo-xylanase (GbtXyl43B). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd08980 | GH43_LbAraf43-like | 1.52e-40 | 892 | 1137 | 2 | 274 | Glycosyl hydrolase family 43 proteins such as Lactobacillus brevis alpha-L-arabinofuranosidase LbAraf43 and Geobacillus thermoleovorans GbtXyl43B. This glycosyl hydrolase family 43 (GH43) subgroup includes enzymes with beta-xylosidase (EC 3.2.1.37), alpha-L-arabinofuranosidase (EC 3.2.1.55) and possibly bifunctional xylosidase/arabinofuranosidase activities. In addition to Lactobacillus brevis alpha-L-arabinofuranosidase LbAraf43 and Geobacillus thermoleovorans IT-08 beta-xylosidase / exo-xylanase (GbtXyl43B). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) familiesGH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18817 | GH43f_LbAraf43-like | 3.20e-40 | 892 | 1139 | 2 | 262 | Glycosyl hydrolase family 43 such as Lactobacillus brevis alpha-L-arabinofuranosidase LbAraf43. This glycosyl hydrolase family 43 (GH43) subgroup includes characterized enzymes with alpha-L-arabinofuranosidase (EC 3.2.1.55) activity. It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. Characterized enzymes belonging to this subgroup include Lactobacillus brevis (LbAraf43) and Weissella sp (WAraf43) which show activity with similar catalytic efficiency on 1,5-alpha-L-arabinooligosaccharides with a degree of polymerization (DP) of 2-3; size is limited by an extended loop at the entrance to the active site. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG3940 | COG3940 | 4.24e-38 | 889 | 1169 | 12 | 313 | Beta-xylosidase, GH43 family [Carbohydrate transport and metabolism]. |
| pfam04616 | Glyco_hydro_43 | 2.87e-30 | 892 | 1164 | 12 | 281 | Glycosyl hydrolases family 43. The glycosyl hydrolase family 43 contains members that are arabinanases. Arabinanases hydrolyze the alpha-1,5-linked L-arabinofuranoside backbone of plant cell wall arabinans. The structure of arabinanase Arb43A from Cellvibrio japonicus reveals a five-bladed beta-propeller fold. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ASK62305.1 | 9.38e-159 | 499 | 1338 | 34 | 722 |
| QJE96570.1 | 2.60e-102 | 881 | 1338 | 24 | 466 |
| QLG40883.1 | 8.91e-52 | 883 | 1339 | 36 | 466 |
| AIQ73224.1 | 1.56e-51 | 883 | 1336 | 35 | 462 |
| AYB42892.1 | 1.80e-50 | 883 | 1339 | 35 | 465 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6QE7_A | 1.15e-29 | 394 | 638 | 52 | 289 | ChainA, Anti-sigma-I factor RsgI3 [Acetivibrio thermocellus],6QE7_B Chain B, Anti-sigma-I factor RsgI3 [Acetivibrio thermocellus],6QE7_C Chain C, Anti-sigma-I factor RsgI3 [Acetivibrio thermocellus] |
| 5M8B_A | 9.39e-29 | 878 | 1175 | 23 | 345 | ChainA, Alpha-L-arabinofuranosidase II [Levilactobacillus brevis],5M8B_B Chain B, Alpha-L-arabinofuranosidase II [Levilactobacillus brevis] |
| 6QDI_A | 1.01e-26 | 394 | 639 | 51 | 294 | ChainA, PA14 domain-containing protein [Acetivibrio clariflavus] |
| 3AKF_A | 7.43e-26 | 892 | 1171 | 21 | 317 | Crystalstructure of exo-1,5-alpha-L-arabinofuranosidase [Streptomyces avermitilis MA-4680 = NBRC 14893],3AKG_A Crystal structure of exo-1,5-alpha-L-arabinofuranosidase complexed with alpha-1,5-L-arabinofuranobiose [Streptomyces avermitilis MA-4680 = NBRC 14893],3AKH_A Crystal structure of exo-1,5-alpha-L-arabinofuranosidase complexed with alpha-1,5-L-arabinofuranotriose [Streptomyces avermitilis MA-4680 = NBRC 14893],3AKI_A Crystal structure of exo-1,5-alpha-L-arabinofuranosidase complexed with alpha-L-arabinofuranosyl azido [Streptomyces avermitilis MA-4680 = NBRC 14893] |
| 5M8E_A | 3.31e-24 | 884 | 1175 | 27 | 343 | Crystalstructure of a GH43 arabonofuranosidase from Weissella sp. strain 142 [Weissella cibaria],5M8E_B Crystal structure of a GH43 arabonofuranosidase from Weissella sp. strain 142 [Weissella cibaria] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DC75 | 7.96e-27 | 394 | 638 | 405 | 642 | Anti-sigma-I factor RsgI3 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=rsgI3 PE=1 SV=1 |
| Q82P90 | 4.73e-25 | 892 | 1171 | 47 | 343 | Extracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Streptomyces avermitilis (strain ATCC 31267 / DSM 46492 / JCM 5070 / NBRC 14893 / NCIMB 12804 / NRRL 8165 / MA-4680) OX=227882 GN=Araf43A PE=1 SV=1 |
| P82594 | 2.47e-23 | 889 | 1141 | 57 | 321 | Extracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Streptomyces chartreusis OX=1969 PE=1 SV=1 |
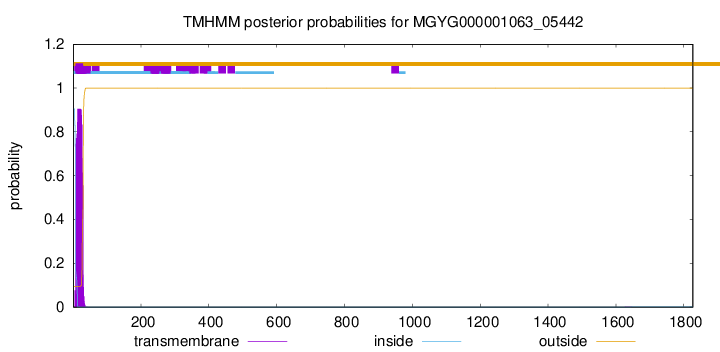
SignalP and Lipop Annotations help
This protein is predicted as SP
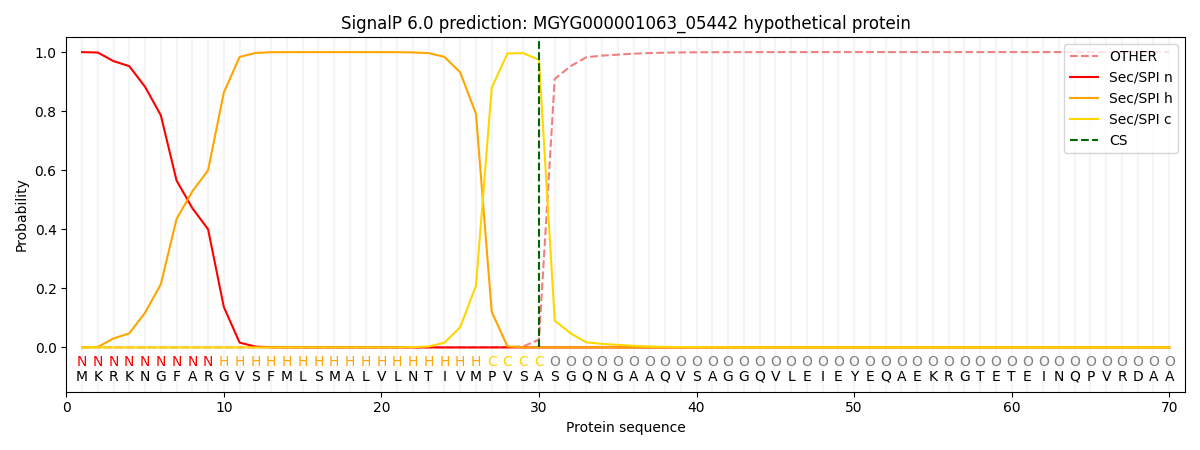
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000384 | 0.998864 | 0.000247 | 0.000170 | 0.000162 | 0.000149 |