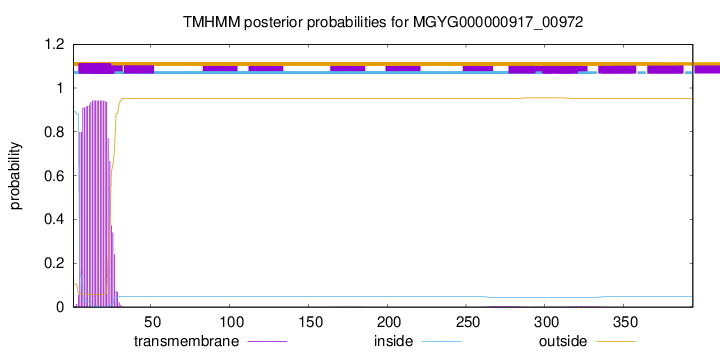You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000917_00972
You are here: Home > Sequence: MGYG000000917_00972
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phascolarctobacterium sp900551745 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Acidaminococcales; Acidaminococcaceae; Phascolarctobacterium; Phascolarctobacterium sp900551745 | |||||||||||
| CAZyme ID | MGYG000000917_00972 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4303; End: 5487 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 114 | 333 | 3.3e-49 | 0.9675925925925926 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 3.07e-76 | 51 | 393 | 1 | 333 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 4.64e-75 | 52 | 370 | 1 | 315 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 2.82e-60 | 60 | 336 | 3 | 282 | beta-hexosaminidase; Provisional |
| PRK15098 | PRK15098 | 3.84e-12 | 42 | 371 | 35 | 351 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBG62799.1 | 1.69e-278 | 1 | 394 | 1 | 394 |
| QNP77560.1 | 1.39e-277 | 1 | 394 | 1 | 394 |
| CBL06667.1 | 2.44e-109 | 21 | 376 | 27 | 383 |
| SNV01291.1 | 5.05e-109 | 21 | 376 | 26 | 384 |
| BDA09432.1 | 5.57e-108 | 21 | 376 | 27 | 383 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6K5J_A | 7.55e-67 | 51 | 380 | 11 | 345 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 4ZM6_A | 1.72e-47 | 52 | 372 | 8 | 337 | Aunique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432],4ZM6_B A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432] |
| 3TEV_A | 6.57e-42 | 58 | 359 | 19 | 318 | Thecrystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1],3TEV_B The crystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1] |
| 3BMX_A | 5.13e-40 | 13 | 373 | 5 | 394 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 4GVF_A | 9.44e-39 | 61 | 357 | 4 | 301 | Crystalstructure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) bound to GlcNAc [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4GVF_B Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) bound to GlcNAc [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4GVG_A Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4GVG_B Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4GVH_A Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) covalently bound to 5-fluoro-GlcNAc. [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4GVH_B Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) covalently bound to 5-fluoro-GlcNAc. [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4HZM_A Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) bound to N-[(3S,4R,5R,6R)-4,5-dihydroxy-6-(hydroxymethyl)piperidin-3-yl]butanamide [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],4HZM_B Crystal structure of Salmonella typhimurium family 3 glycoside hydrolase (NagZ) bound to N-[(3S,4R,5R,6R)-4,5-dihydroxy-6-(hydroxymethyl)piperidin-3-yl]butanamide [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B0BQ51 | 2.08e-43 | 60 | 347 | 1 | 291 | Beta-hexosaminidase OS=Actinobacillus pleuropneumoniae serotype 3 (strain JL03) OX=434271 GN=nagZ PE=3 SV=1 |
| A3N1B7 | 2.91e-43 | 60 | 347 | 1 | 291 | Beta-hexosaminidase OS=Actinobacillus pleuropneumoniae serotype 5b (strain L20) OX=416269 GN=nagZ PE=3 SV=1 |
| B3GXZ7 | 2.91e-43 | 60 | 347 | 1 | 291 | Beta-hexosaminidase OS=Actinobacillus pleuropneumoniae serotype 7 (strain AP76) OX=537457 GN=nagZ PE=3 SV=1 |
| B2FPW9 | 1.23e-42 | 60 | 361 | 1 | 300 | Beta-hexosaminidase OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=nagZ PE=3 SV=1 |
| C5BFM6 | 1.93e-42 | 61 | 336 | 4 | 281 | Beta-hexosaminidase OS=Edwardsiella ictaluri (strain 93-146) OX=634503 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000003 | 1.000040 | 0.000000 | 0.000000 | 0.000000 |