You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004876_00662
You are here: Home > Sequence: MGYG000004876_00662
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; | |||||||||||
| CAZyme ID | MGYG000004876_00662 | |||||||||||
| CAZy Family | PL8 | |||||||||||
| CAZyme Description | Chondroitinase-AC | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 96765; End: 98945 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL8 | 365 | 611 | 8.8e-74 | 0.9879518072289156 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01083 | GAG_Lyase | 1.83e-147 | 80 | 689 | 38 | 693 | Glycosaminoglycan (GAG) polysaccharide lyase family. This family consists of a group of secreted bacterial lyase enzymes capable of acting on glycosaminoglycans, such as hyaluronan and chondroitin, in the extracellular matrix of host tissues, contributing to the invasive capacity of the pathogen. These are broad-specificity glycosaminoglycan lyases which recognize uronyl residues in polysaccharides and cleave their glycosidic bonds via a beta-elimination reaction to form a double bond between C-4 and C-5 of the non-reducing terminal uronyl residues of released products. Substrates include chondroitin, chondroitin 4-sulfate, chondroitin 6-sulfate, and hyaluronic acid. Family members include chondroitin AC lyase, chondroitin abc lyase, xanthan lyase, and hyalurate lyase. |
| pfam02278 | Lyase_8 | 1.62e-89 | 365 | 614 | 1 | 252 | Polysaccharide lyase family 8, super-sandwich domain. This family consists of a group of secreted bacterial lyase enzymes EC:4.2.2.1 capable of acting on hyaluronan and chondroitin in the extracellular matrix of host tissues, contributing to the invasive capacity of the pathogen. |
| pfam08124 | Lyase_8_N | 4.49e-29 | 78 | 313 | 31 | 282 | Polysaccharide lyase family 8, N terminal alpha-helical domain. This family consists of a group of secreted bacterial lyase enzymes EC:4.2.2.1 capable of acting on hyaluronan and chondroitin in the extracellular matrix of host tissues, contributing to the invasive capacity of the pathogen. |
| pfam02884 | Lyase_8_C | 1.41e-04 | 629 | 691 | 1 | 61 | Polysaccharide lyase family 8, C-terminal beta-sandwich domain. This family consists of a group of secreted bacterial lyase enzymes EC:4.2.2.1 capable of acting on hyaluronan and chondroitin in the extracellular matrix of host tissues, contributing to the invasive capacity of the pathogen. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ58447.1 | 0.0 | 1 | 721 | 1 | 721 |
| QUT90440.1 | 0.0 | 1 | 721 | 1 | 721 |
| QDO71119.1 | 0.0 | 1 | 721 | 1 | 721 |
| QUT47778.1 | 2.24e-263 | 26 | 725 | 27 | 724 |
| QIK58581.1 | 7.39e-231 | 8 | 716 | 7 | 713 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1CB8_A | 1.02e-79 | 84 | 718 | 32 | 674 | CHONDROITINASEAC LYASE FROM FLAVOBACTERIUM HEPARINUM [Pedobacter heparinus] |
| 1HM2_A | 1.62e-79 | 84 | 718 | 54 | 696 | ACTIVESITE OF CHONDROITINASE AC LYASE REVEALED BY THE STRUCTURE OF ENZYME-OLIGOSACCHARIDE COMPLEXES AND MUTAGENESIS [Pedobacter heparinus],1HM3_A Active Site Of Chondroitinase Ac Lyase Revealed By The Structure Of Enzyme-Oligosaccharide Complexes And Mutagenesis [Pedobacter heparinus],1HMU_A ACTIVE SITE OF CHONDROITINASE AC LYASE REVEALED BY THE STRUCTURE OF ENZYME-OLIGOSACCHARIDE COMPLEXES AND MUTAGENESIS [Pedobacter heparinus],1HMW_A Active Site Of Chondroitinase Ac Lyase Revealed By The Structure Of Enzyme-oligosaccharide Complexes And Mutagenesis [Pedobacter heparinus] |
| 1RW9_A | 1.59e-37 | 155 | 692 | 124 | 706 | Crystalstructure of the Arthrobacter aurescens chondroitin AC lyase [Paenarthrobacter aurescens] |
| 1RWA_A | 1.59e-37 | 155 | 692 | 124 | 706 | Crystalstructure of Arthrobacter aurescens chondroitin AC lyase [Paenarthrobacter aurescens],1RWC_A Crystal structure of Arthrobacter aurescens chondroitin AC lyase [Paenarthrobacter aurescens],1RWF_A Crystal structure of Arthrobacter aurescens chondroitin AC lyase in complex with chondroitin tetrasaccharide [Paenarthrobacter aurescens],1RWG_A Crystal structure of Arthrobacter aurescens chondroitin AC lyase in complex with chondroitin tetrasaccharide [Paenarthrobacter aurescens],1RWH_A Crystal structure of Arthrobacter aurescens chondroitin AC lyase in complex with chondroitin tetrasaccharide [Paenarthrobacter aurescens] |
| 1J0M_A | 6.49e-33 | 113 | 694 | 73 | 702 | CrystalStructure of Bacillus sp. GL1 Xanthan Lyase that Acts on Side Chains of Xanthan [Bacillus sp. GL1],1J0N_A Crystal Structure of Bacillus sp. GL1 Xanthan Lyase that Acts on Side Chains of Xanthan [Bacillus sp. GL1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q59288 | 8.86e-79 | 84 | 718 | 54 | 696 | Chondroitinase-AC OS=Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / CIP 104194 / JCM 7457 / NBRC 12017 / NCIMB 9290 / NRRL B-14731 / HIM 762-3) OX=485917 GN=cslA PE=1 SV=1 |
| Q9AQS0 | 6.99e-32 | 113 | 694 | 98 | 727 | Xanthan lyase OS=Bacillus sp. (strain GL1) OX=84635 GN=xly PE=1 SV=1 |
| Q54873 | 1.76e-22 | 79 | 718 | 325 | 1024 | Hyaluronate lyase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=SP_0314 PE=1 SV=2 |
| Q59801 | 1.67e-21 | 148 | 699 | 175 | 778 | Hyaluronate lyase OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=hysA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
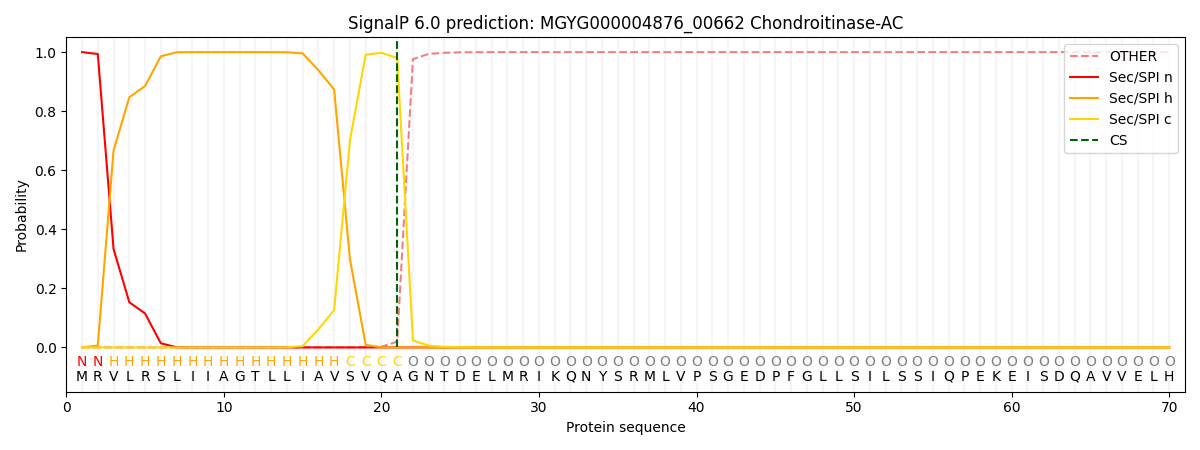
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000474 | 0.998651 | 0.000266 | 0.000198 | 0.000194 | 0.000179 |
