You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004847_00016
You are here: Home > Sequence: MGYG000004847_00016
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1670 sp902406135 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Anaerotignaceae; UMGS1670; UMGS1670 sp902406135 | |||||||||||
| CAZyme ID | MGYG000004847_00016 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12010; End: 13665 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 209 | 531 | 4e-37 | 0.8074324324324325 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd02874 | GH18_CFLE_spore_hydrolase | 7.35e-38 | 256 | 532 | 43 | 304 | Cortical fragment-lytic enzyme (CFLE) is a peptidoglycan hydrolase involved in bacterial endospore germination. CFLE is expressed as an inactive preprotein (called SleB) in the forespore compartment of sporulating cells. SleB translocates across the forespore inner membrane and is deposited as a mature enzyme in the cortex layer of the spore. As part of a sensory mechanism capable of initiating germination, CFLE degrades a spore-specific peptidoglycan constituent called muramic-acid delta-lactam that comprises the outer cortex. CFLE has a C-terminal glycosyl hydrolase family 18 (GH18) catalytic domain as well as two N-terminal LysM peptidoglycan-binding domains. In addition to SleB, this family includes YaaH, YdhD, and YvbX from Bacillus subtilis. |
| COG3858 | YaaH | 4.45e-28 | 274 | 532 | 168 | 409 | Spore germination protein YaaH [Cell cycle control, cell division, chromosome partitioning]. |
| smart00636 | Glyco_18 | 3.93e-23 | 284 | 532 | 78 | 333 | Glyco_18 domain. |
| pfam00704 | Glyco_hydro_18 | 6.95e-22 | 282 | 531 | 73 | 305 | Glycosyl hydrolases family 18. |
| cd06549 | GH18_trifunctional | 3.30e-18 | 210 | 531 | 12 | 291 | GH18 domain of an uncharacterized family of bacterial proteins, which share a common three-domain architecture: an N-terminal glycosyl hydrolase family 18 (GH18) domain, a glycosyl transferase family 2 domain, and a C-terminal polysaccharide deacetylase domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QOX64072.1 | 1.04e-165 | 7 | 535 | 13 | 549 |
| QHI71217.1 | 6.01e-154 | 2 | 538 | 5 | 546 |
| QIB69198.1 | 6.01e-154 | 19 | 538 | 25 | 546 |
| QAT42616.1 | 1.20e-153 | 19 | 538 | 25 | 546 |
| QGG48974.1 | 1.21e-146 | 27 | 551 | 35 | 581 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4WIW_A | 1.78e-13 | 299 | 536 | 111 | 335 | ChainA, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2],4WIW_B Chain B, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2],4WIW_C Chain C, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2],4WIW_D Chain D, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2],4WIW_E Chain E, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2],4WIW_F Chain F, Glycoside hydrolase family 18 [Desulfitobacterium hafniense DCB-2] |
| 4Q6T_A | 3.31e-10 | 321 | 551 | 114 | 340 | Thecrystal structure of a class V chitininase from Pseudomonas fluorescens Pf-5 [Pseudomonas protegens Pf-5] |
| 4S3J_A | 4.40e-09 | 324 | 532 | 215 | 412 | Crystalstructure of the Bacillus cereus spore cortex-lytic enzyme SleL [Bacillus cereus ATCC 10876],4S3J_B Crystal structure of the Bacillus cereus spore cortex-lytic enzyme SleL [Bacillus cereus ATCC 10876],4S3J_C Crystal structure of the Bacillus cereus spore cortex-lytic enzyme SleL [Bacillus cereus ATCC 10876] |
| 5GZU_A | 5.26e-09 | 335 | 528 | 659 | 863 | CrystalStructure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery [Paenibacillus sp. FPU-7],5GZU_B Crystal Structure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery [Paenibacillus sp. FPU-7],5GZV_A Crystal Structure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery [Paenibacillus sp. FPU-7],5GZV_B Crystal Structure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery [Paenibacillus sp. FPU-7] |
| 5GZT_B | 5.67e-09 | 335 | 528 | 910 | 1114 | CrystalStructure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery [Paenibacillus sp. FPU-7] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P38535 | 6.65e-17 | 24 | 201 | 905 | 1086 | Exoglucanase XynX OS=Acetivibrio thermocellus OX=1515 GN=xynX PE=3 SV=1 |
| P38536 | 4.19e-16 | 27 | 198 | 1682 | 1857 | Amylopullulanase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=amyB PE=3 SV=2 |
| P19424 | 1.06e-12 | 24 | 192 | 38 | 210 | Endoglucanase OS=Bacillus sp. (strain KSM-635) OX=1415 PE=1 SV=1 |
| C6CRV0 | 1.23e-12 | 27 | 194 | 1283 | 1457 | Endo-1,4-beta-xylanase A OS=Paenibacillus sp. (strain JDR-2) OX=324057 GN=xynA1 PE=1 SV=1 |
| P36917 | 5.98e-09 | 32 | 126 | 1060 | 1155 | Endo-1,4-beta-xylanase A OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
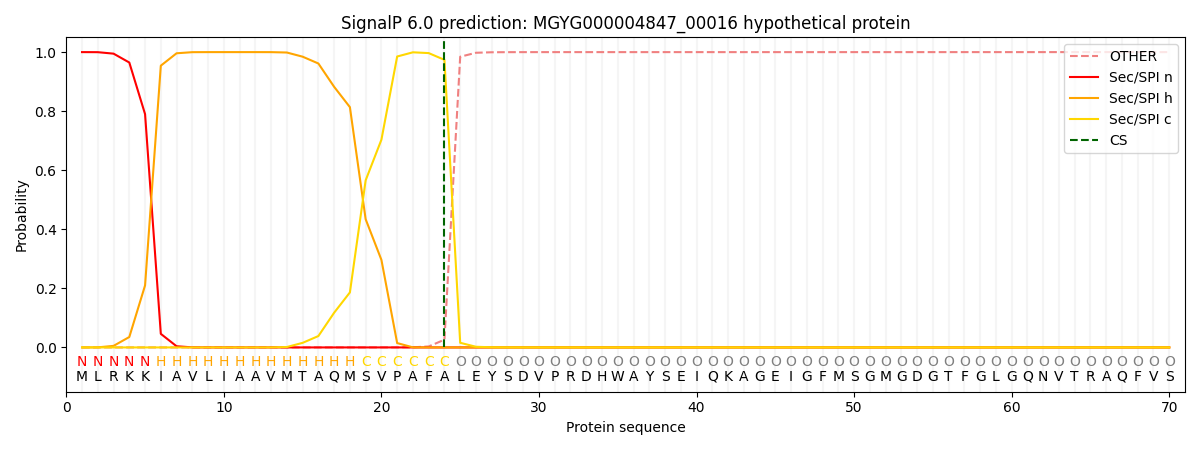
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000254 | 0.999007 | 0.000187 | 0.000185 | 0.000172 | 0.000149 |
