You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004820_00670
You are here: Home > Sequence: MGYG000004820_00670
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; UBA7173; | |||||||||||
| CAZyme ID | MGYG000004820_00670 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5206; End: 6252 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 47 | 332 | 8e-85 | 0.9936507936507937 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd08991 | GH43_HoAraf43-like | 6.76e-114 | 49 | 340 | 1 | 283 | Glycosyl hydrolase family 43 protein such as Halothermothrix orenii H 168 alpha-L-arabinofuranosidase (HoAraf43;Hore_20580). This glycosyl hydrolase family 43 (GH43) subgroup includes Halothermothrix orenii H 168 alpha-L-arabinofuranosidase (EC 3.2.1.55) (HoAraf43;Hore_20580). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. This GH43_ HoAraf43-like subgroup includes enzymes that have been annotated as having xylan-digesting beta-xylosidase (EC 3.2.1.37) and xylanase (endo-alpha-L-arabinanase, EC 3.2.1.8) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| pfam04616 | Glyco_hydro_43 | 1.32e-61 | 34 | 332 | 1 | 279 | Glycosyl hydrolases family 43. The glycosyl hydrolase family 43 contains members that are arabinanases. Arabinanases hydrolyze the alpha-1,5-linked L-arabinofuranoside backbone of plant cell wall arabinans. The structure of arabinanase Arb43A from Cellvibrio japonicus reveals a five-bladed beta-propeller fold. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd08999 | GH43_ABN-like | 9.03e-59 | 49 | 330 | 9 | 280 | Glycosyl hydrolase family 43 protein such as endo-alpha-L-arabinanase. This glycosyl hydrolase family 43 (GH43) subgroup includes mostly enzymes with alpha-L-arabinofuranosidase (ABF; EC 3.2.1.55) and endo-alpha-L-arabinanase (ABN; EC 3.2.1.99) activities. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. The GH43 ABN enzymes hydrolyze alpha-1,5-L-arabinofuranoside linkages while the ABF enzymes cleave arabinose side chains so that the combined actions of these two enzymes reduce arabinan to L-arabinose and/or arabinooligosaccharides. These arabinan-degrading enzymes are important in the food industry for efficient production of L-arabinose from agricultural waste; L-arabinose is often used as a bioactive sweetener. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd09004 | GH43_bXyl-like | 2.50e-58 | 49 | 335 | 1 | 264 | Glycosyl hydrolase family 43 protein such as Bacteroides thetaiotaomicron VPI-5482 alpha-L-arabinofuranosidases (BT3675;BT_3675) and (BT3662;BT_3662); includes mostly xylanases. This glycosyl hydrolase family 43 (GH43) subgroup includes enzymes that have been annotated as xylan-digesting beta-xylosidase (EC 3.2.1.37) and xylanase (endo-alpha-L-arabinanase, EC 3.2.1.8) activities, as well the Bacteroides thetaiotaomicron VPI-5482 alpha-L-arabinofuranosidases (EC 3.2.1.55) (BT3675;BT_3675) and (BT3662;BT_3662). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd08978 | GH_F | 3.46e-55 | 49 | 307 | 1 | 251 | Glycosyl hydrolase families 43 and 62 form CAZY clan GH-F. This glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) includes family 43 (GH43) and 62 (GH62). GH43 includes enzymes with beta-xylosidase (EC 3.2.1.37), beta-1,3-xylosidase (EC 3.2.1.-), alpha-L-arabinofuranosidase (EC 3.2.1.55), arabinanase (EC 3.2.1.99), xylanase (EC 3.2.1.8), endo-alpha-L-arabinanases (beta-xylanases) and galactan 1,3-beta-galactosidase (EC 3.2.1.145) activities. GH62 includes enzymes characterized as arabinofuranosidases (alpha-L-arabinofuranosidases; EC 3.2.1.55) that specifically cleave either alpha-1,2 or alpha-1,3-L-arabinofuranose side chains from xylans. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many of the enzymes in this family display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. GH62 are also predicted to be inverting enzymes. A common structural feature of both, GH43 and GH62 enzymes, is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGA24609.1 | 5.50e-163 | 33 | 347 | 31 | 344 |
| BCG53971.1 | 3.88e-147 | 46 | 347 | 2 | 303 |
| SCM59045.1 | 1.10e-128 | 14 | 339 | 8 | 332 |
| AGB41264.1 | 1.74e-86 | 28 | 347 | 38 | 351 |
| ACL70803.1 | 1.36e-84 | 33 | 347 | 4 | 312 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4QQS_A | 4.98e-86 | 33 | 347 | 4 | 312 | Crystalstructure of a thermostable family-43 glycoside hydrolase [Halothermothrix orenii H 168],4QQS_B Crystal structure of a thermostable family-43 glycoside hydrolase [Halothermothrix orenii H 168] |
| 3KST_A | 6.07e-41 | 40 | 344 | 16 | 303 | Crystalstructure of Endo-1,4-beta-xylanase (NP_811807.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.70 A resolution [Bacteroides thetaiotaomicron VPI-5482],3KST_B Crystal structure of Endo-1,4-beta-xylanase (NP_811807.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.70 A resolution [Bacteroides thetaiotaomicron VPI-5482] |
| 3QZ4_A | 3.15e-25 | 49 | 340 | 16 | 310 | Crystalstructure of an Endo-1,4-beta-xylanase D (BT_3675) from Bacteroides thetaiotaomicron VPI-5482 at 1.74 A resolution [Bacteroides thetaiotaomicron],3QZ4_B Crystal structure of an Endo-1,4-beta-xylanase D (BT_3675) from Bacteroides thetaiotaomicron VPI-5482 at 1.74 A resolution [Bacteroides thetaiotaomicron] |
| 5MSX_A | 2.19e-21 | 48 | 335 | 34 | 302 | Glycosidehydrolase BT_3662 [Bacteroides thetaiotaomicron VPI-5482],5MSX_B Glycoside hydrolase BT_3662 [Bacteroides thetaiotaomicron VPI-5482],5MSX_C Glycoside hydrolase BT_3662 [Bacteroides thetaiotaomicron VPI-5482] |
| 6MS3_A | 1.08e-19 | 18 | 332 | 14 | 311 | Crystalstructure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580],6MS3_B Crystal structure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q82P90 | 3.54e-16 | 7 | 332 | 4 | 334 | Extracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Streptomyces avermitilis (strain ATCC 31267 / DSM 46492 / JCM 5070 / NBRC 14893 / NCIMB 12804 / NRRL 8165 / MA-4680) OX=227882 GN=Araf43A PE=1 SV=1 |
| P82594 | 1.40e-11 | 49 | 305 | 59 | 317 | Extracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Streptomyces chartreusis OX=1969 PE=1 SV=1 |
| P48791 | 1.11e-09 | 103 | 335 | 84 | 316 | Beta-xylosidase OS=Prevotella ruminicola OX=839 GN=xynB PE=3 SV=1 |
| P49943 | 2.87e-08 | 41 | 334 | 9 | 317 | Xylosidase/arabinosidase OS=Bacteroides ovatus OX=28116 GN=xsa PE=2 SV=1 |
| B3EYM8 | 1.17e-07 | 40 | 332 | 18 | 309 | Intracellular endo-alpha-(1->5)-L-arabinanase OS=Geobacillus stearothermophilus OX=1422 GN=abnB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
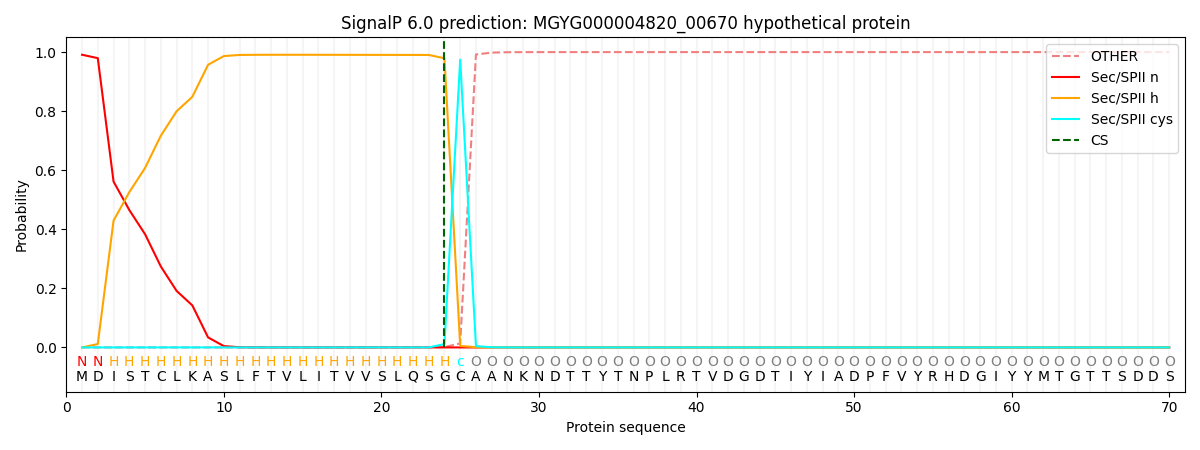
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000017 | 0.008663 | 0.991345 | 0.000004 | 0.000006 | 0.000004 |
