You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004697_00862
You are here: Home > Sequence: MGYG000004697_00862
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
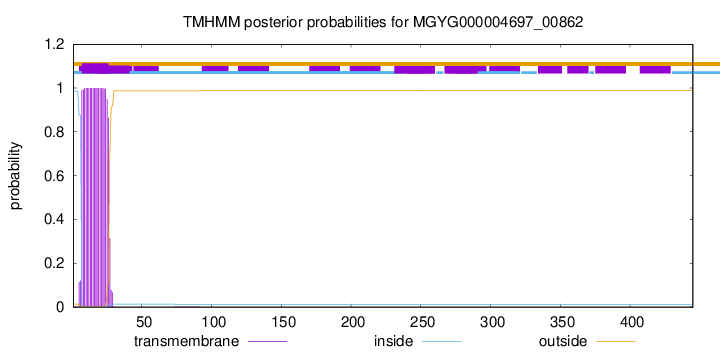
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eubacterium_F; | |||||||||||
| CAZyme ID | MGYG000004697_00862 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | Endo-1,4-beta-xylanase Y | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 75300; End: 76637 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 68 | 435 | 8.4e-85 | 0.9867986798679867 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 3.00e-91 | 113 | 433 | 1 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 8.69e-86 | 68 | 433 | 2 | 308 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 4.92e-64 | 72 | 431 | 31 | 335 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCS82260.1 | 1.88e-165 | 63 | 439 | 66 | 419 |
| AZT91389.1 | 4.37e-164 | 63 | 439 | 66 | 419 |
| ADQ03735.1 | 6.04e-160 | 57 | 444 | 99 | 466 |
| AAA23062.1 | 3.55e-142 | 113 | 439 | 1 | 304 |
| ABP67982.1 | 3.55e-142 | 113 | 439 | 1 | 304 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2W5F_A | 5.36e-54 | 68 | 430 | 185 | 521 | ChainA, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2W5F_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus] |
| 2WYS_A | 2.82e-51 | 68 | 430 | 185 | 521 | ChainA, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WYS_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WZE_A Chain A, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WZE_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus] |
| 6D5C_A | 5.71e-49 | 71 | 435 | 27 | 349 | Structureof Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_B Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_C Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii] |
| 5Y3X_A | 2.58e-48 | 71 | 434 | 36 | 355 | Crystalstructure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_B Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_C Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_D Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_E Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL],5Y3X_F Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis [Caldicellulosiruptor owensensis OL] |
| 4L4O_A | 1.44e-47 | 71 | 434 | 17 | 335 | Thecrystal structure of CbXyn10B in native form [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23557 | 7.10e-143 | 113 | 439 | 1 | 304 | Putative endo-1,4-beta-xylanase OS=Caldicellulosiruptor saccharolyticus OX=44001 PE=3 SV=1 |
| P26223 | 3.62e-64 | 68 | 438 | 4 | 339 | Endo-1,4-beta-xylanase B OS=Butyrivibrio fibrisolvens OX=831 GN=xynB PE=3 SV=1 |
| Q60042 | 4.11e-52 | 63 | 438 | 361 | 690 | Endo-1,4-beta-xylanase A OS=Thermotoga neapolitana OX=2337 GN=xynA PE=1 SV=1 |
| P51584 | 4.26e-52 | 68 | 430 | 196 | 532 | Endo-1,4-beta-xylanase Y OS=Acetivibrio thermocellus OX=1515 GN=xynY PE=1 SV=1 |
| P23556 | 4.31e-51 | 71 | 435 | 21 | 341 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
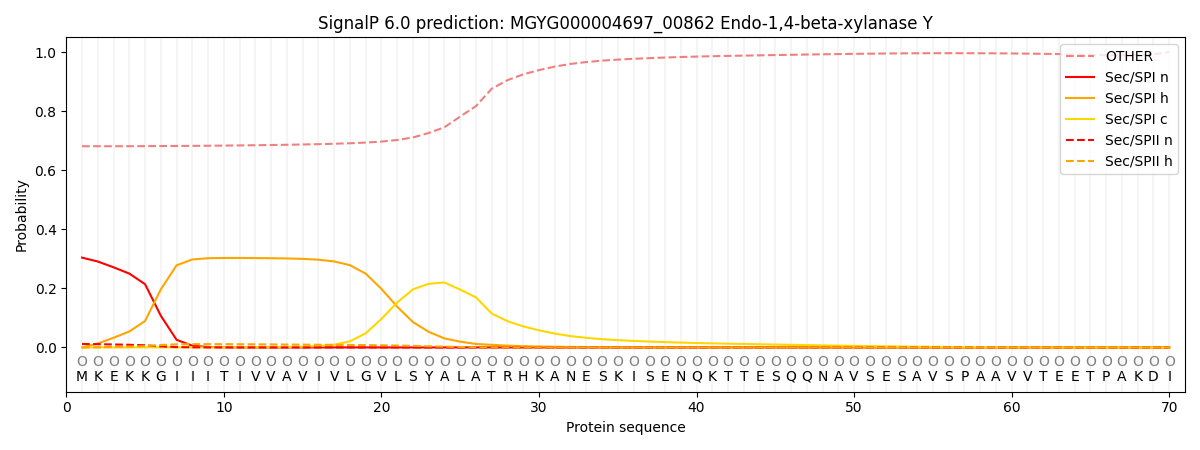
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.695915 | 0.280812 | 0.013888 | 0.002651 | 0.001133 | 0.005588 |